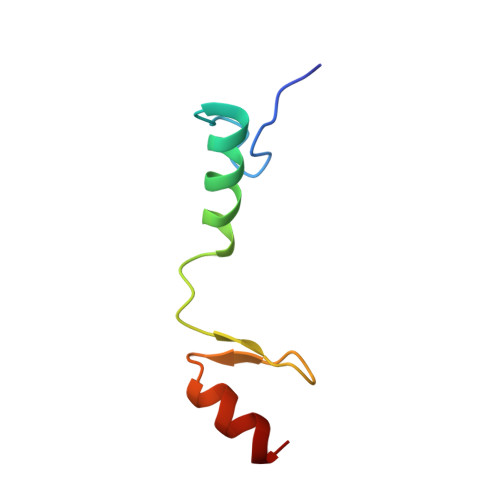
Solution NMR Structure of Zinc finger protein 423 from Homo sapiens, Northeast Structural Genomics Consortium (NESG) Target HR7298F
Pederson, K., Lee, D., Kohan, E., Janjua, H., Xiao, R., Everett, J.K., Acton, T.B., Montelione, G.T., Prestegard, J.H.To be published.