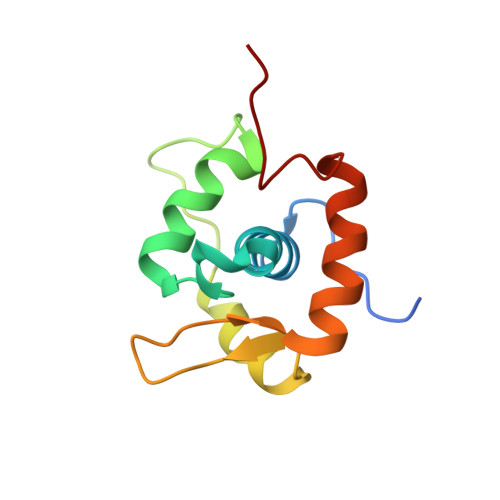
A winged helix domain in human MUS81 binds DNA and modulates the endonuclease activity of MUS81 complexes.
Fadden, A.J., Schalbetter, S., Bowles, M., Harris, R., Lally, J., Carr, A.M., McDonald, N.Q.(2013) Nucleic Acids Res 41: 9741-9752
- PubMed: 23982516 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gkt760
- Primary Citation Related Structures:
2MC3 - PubMed Abstract:
The MUS81-EME1 endonuclease maintains metazoan genomic integrity by cleaving branched DNA structures that arise during the resolution of recombination intermediates. In humans, MUS81 also forms a poorly characterized complex with EME2. Here, we identify and determine the structure of a winged helix (WH) domain from human MUS81, which binds DNA. WH domain mutations greatly reduce binding of the isolated domain to DNA and impact on incision activity of MUS81-EME1/EME2 complexes. Deletion of the WH domain reduces the endonuclease activity of both MUS81-EME1 and MUS81-EME2 complexes, and incisions made by MUS81-EME2 are made closer to the junction on substrates containing a downstream duplex, such as fork structures and nicked Holliday junctions. WH domain mutation or deletion in Schizosaccharomyces pombe phenocopies the DNA-damage sensitivity of strains deleted for mus81. Our results indicate an important role for the WH domain in both yeast and human MUS81 complexes.
- Structural Biology Laboratory, Cancer Research UK, London Research Institute, 44 Lincoln's Inn Fields, London, WC2A 3LY, UK, Genome Damage and Stability Centre, University of Sussex, Brighton, BN1 9RQ, UK and Institute of Structural and Molecular Biology, University College London and Birkbeck College, Malet Street, London WC1E 7HX, UK.
Organizational Affiliation: