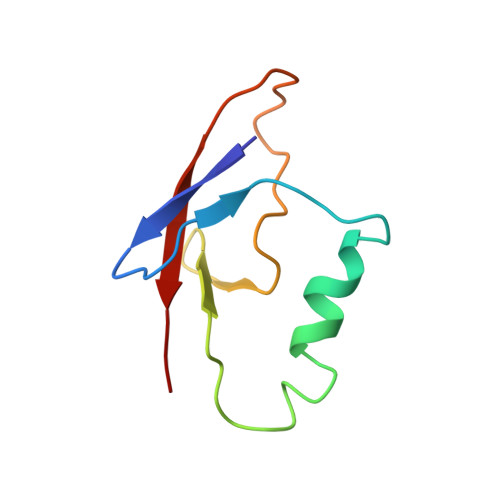
Disruption of FAT10-MAD2 binding inhibits tumor progression.
Theng, S.S., Wang, W., Mah, W.C., Chan, C., Zhuo, J., Gao, Y., Qin, H., Lim, L., Chong, S.S., Song, J., Lee, C.G.(2014) Proc Natl Acad Sci U S A 111: E5282-E5291
- PubMed: 25422469
- DOI: https://doi.org/10.1073/pnas.1403383111
- Primary Citation Related Structures:
2MBE - PubMed Abstract:
FAT10 (HLA-F-adjacent transcript 10) is a ubiquitin-like modifier that is commonly overexpressed in various tumors. It was found to play a role in mitotic regulation through its interaction with mitotic arrest-deficient 2 (MAD2). Overexpression of FAT10 promotes tumor growth and malignancy. Here, we identified the MAD2-binding interface of FAT10 to be located on its first ubiquitin-like domain whose NMR structure thus was determined. We further proceeded to demonstrate that disruption of the FAT10-MAD2 interaction through mutation of specific MAD2-binding residues did not interfere with the interaction of FAT10 with its other known interacting partners. Significantly, ablation of the FAT10-MAD2 interaction dramatically limited the promalignant capacity of FAT10, including promoting tumor growth in vivo and inducing aneuploidy, proliferation, migration, invasion, and resistance to apoptosis in vitro. Our results strongly suggest that the interaction of FAT10 with MAD2 is a key mechanism underlying the promalignant property of FAT10 and offer prospects for the development of anticancer strategies.
- Department of Biochemistry, Yong Loo Lin School of Medicine, National University of Singapore, Singapore 119077, Singapore; National University of Singapore Graduate School of Integrative Sciences and Engineering, National University of Singapore, Singapore 119077, Singapore;
Organizational Affiliation: