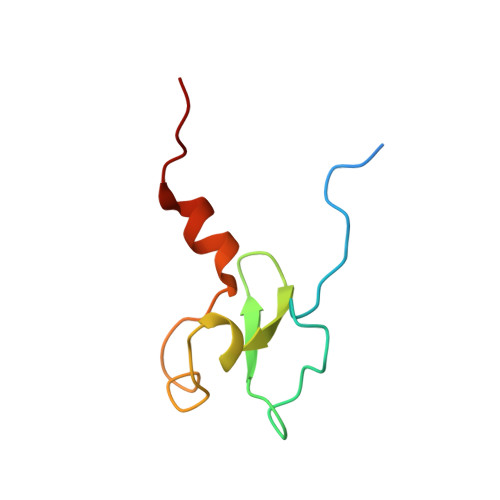
PHD domain from human SHPRH.
Machado, L.E., Pustovalova, Y., Kile, A.C., Pozhidaeva, A., Cimprich, K.A., Almeida, F.C., Bezsonova, I., Korzhnev, D.M.(2013) J Biomol NMR 56: 393-399
- PubMed: 23907177 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1007/s10858-013-9758-2
- Primary Citation Related Structures:
2M85 - PubMed Abstract:
SHPRH ( S NF2, h istone linker, P HD, R ING, h elicase) is a SWI2/SNF2-family ATP-dependent chromatin remodeling factor, and one of E3 ubiquitin ligases responsible for Ubc13-Mms2-dependent K63 poly-ubiquitination of PCNA ( p roliferating c ell n uclear a ntigen) that promotes error-free DNA damage tolerance in eukaryotes. In contrast to its functional homologues, S. cerevisiae Rad5 and human HLTF ( h elicase l ike t ranscription f actor), SHPRH contains a PHD ( p lant h omeo d omain) finger embedded in the ‘minor’ insert region of the core helicase-like domain. PHD fingers are often found in proteins involved in chromatin remodeling and transcription regulation, and are generally considered as ‘readers’ of methylation state of histone tails, primarily the lysine 4 (K4) residue of histone H3 (H3K4). Here we report the solution NMR structure of the SHPRH PHD domain and investigate whether this domain is capable of recognizing H3K4 modifications. The domain adopts a canonical PHD-finger fold with a central two-stranded anti-parallel β-sheet flanked on both sides by the two interleaved zinc-binding sites. Despite the presence of a subset of aromatic residues characteristic for PHD-fingers that preferentially bind methylated H3K4, NMR titration experiments reveal that SHPRH PHD does not specifically interact with the H3-derived peptides irrespective of K4 methylation. This result suggests that the SHPRH PHD domain might have evolved a different function other than recognizing histone modifications.
- Department of Molecular, Microbial and Structural Biology, University of Connecticut Health Center, Farmington, CT 06030, USA.
Organizational Affiliation: