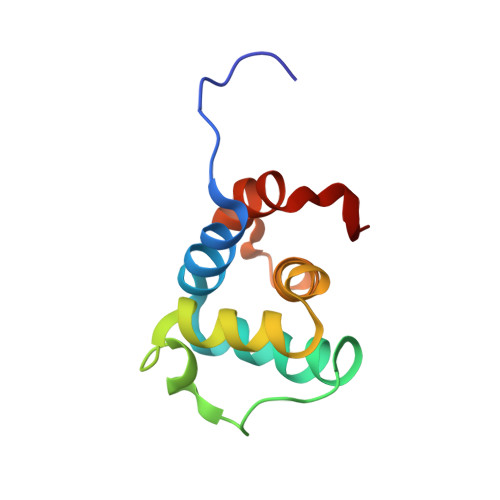
Three-Dimensional Structure of Human NLRP10/PYNOD Pyrin Domain Reveals a Homotypic Interaction Site Distinct from Its Mouse Homologue.
Su, M.Y., Kuo, C.I., Chang, C.F., Chang, C.I.(2013) PLoS One 8: e67843-e67843
- PubMed: 23861819 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pone.0067843
- Primary Citation Related Structures:
2M5V - PubMed Abstract:
NLRPs (Nucleotide-binding domain, leucine-rich repeat and pyrin domain containing proteins) are a family of pattern-recognition receptors (PRRs) that sense intracellular microbial components and endogenous stress signals. NLRP10 (also known as PYNOD) is a unique NLRP member characterized by a lack of the putative ligand-binding leucine-rich repeat domain. Recently, human NLRP10 has been shown to inhibit the self-association of ASC into aggregates and ASC-mediated procaspase-1 processing. However, such activities are not found in mouse NLRP10. Here we report the solution structure and dynamics of human NLRP10 pyrin domain (PYD), whose helix H3 and loop H2-H3 adopt a conformation distinct from those of mouse NLRP10. Docking studies show that human and mouse NLRP10 PYDs may interact differently with ASC PYD. These results provide a possible structural explanation for the contrasting effect of NLRP10 on ASC aggregation in human cells versus mouse models. Finally, we also provide evidence that in human NLRP10 the PYD domain may not interact with the NOD domain to regulate its intrinsic nucleotide hydrolysis activity.
- Institute of Biological Chemistry, Academia Sinica, Taipei, Taiwan, Republic of China.
Organizational Affiliation: