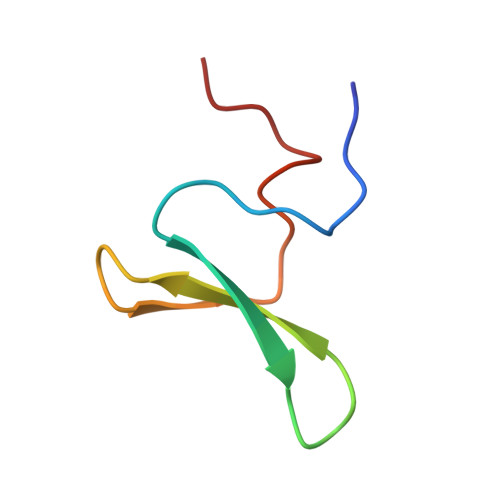
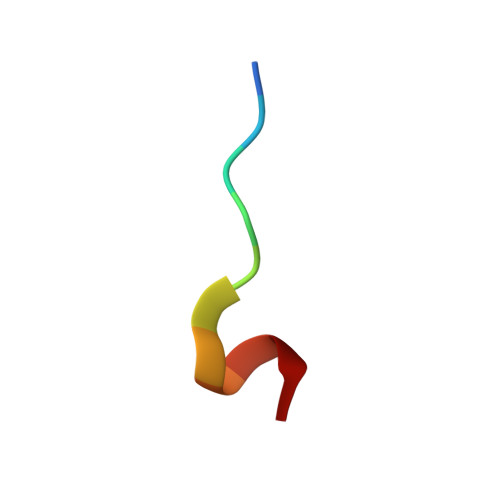
Structure and dynamics of human Nedd4-1 WW3 in complex with the alpha ENaC PY motif.
Bobby, R., Medini, K., Neudecker, P., Lee, T.V., Brimble, M.A., McDonald, F.J., Lott, J.S., Dingley, A.J.(2013) Biochim Biophys Acta 1834: 1632-1641
- PubMed: 23665454 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbapap.2013.04.031
- Primary Citation Related Structures:
2M3O - PubMed Abstract:
Nedd4-1 (neuronal precursor cell expressed developmentally downregulated gene 4-1) is an E3 ubiquitin ligase that interacts with and negatively regulates the epithelial Na(+) channel (ENaC). The WW domains of Nedd4-1 bind to the ENaC subunits via recognition of PY motifs. Human Nedd4-1 (hNedd4-1) contains four WW domains with the third domain (WW3*) showing the strongest affinity to the PY motif. To understand the mechanism underlying this binding affinity, we have carried out NMR structural and dynamics analyses of the hNedd4-1 WW3* domain in complex with a peptide comprising the C-terminal tail of the human ENaC α-subunit. The structure reveals that the peptide interacts in a similar manner to other WW domain-ENaC peptide structures. Crucial interactions that likely provide binding affinity are the broad XP groove facilitating additional contacts between the WW3* domain and the peptide, compared to similar complexes, and the large surface area buried (83Å(2)) between R430 (WW3*) and L647' (αENaC). This corroborates the model-free analysis of the (15)N backbone relaxation data, which showed that R430 is the most rigid residue in the domain (S(2)=0.90±0.01). Carr-Purcell-Meiboom-Gill relaxation dispersion analysis identified two different conformational exchange processes on the μs-ms time-scale. One of these processes involves residues located at the peptide binding interface, suggesting conformational exchange may play a role in peptide recognition. Thus, both structural and dynamic features of the complex appear to define the high binding affinity. The results should aid interpretation of biochemical data and modeling interfaces between Nedd4-1 and other interacting proteins.
- School of Chemical Sciences, The University of Auckland, Auckland, New Zealand.
Organizational Affiliation: