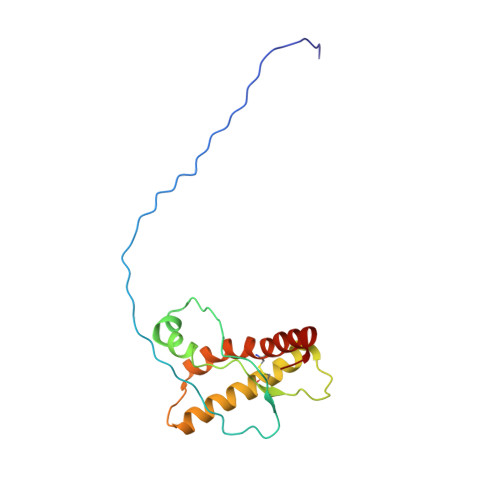
Structural Rearrangements at Physiological pH: Nuclear Magnetic Resonance Insights from the V210I Human Prion Protein Mutant.
Biljan, I., Ilc, G., Giachin, G., Plavec, J., Legname, G.(2012) Biochemistry 51: 7465-7474
- PubMed: 22947063 Search on PubMed
- DOI: https://doi.org/10.1021/bi3009856
- Primary Citation Related Structures:
2LV1 - PubMed Abstract:
A major focus in prion structural biology studies is unraveling the molecular mechanism leading to the structural conversion of PrP(C) to its pathological form, PrP(Sc). In our recent studies, we attempted to understand the early events of the conformational changes leading to PrP(Sc) using as investigative tools point mutations clustered in the open reading frame of the human PrP gene and linked to genetic forms of human prion diseases. In the work presented here, we investigate the effect of pH on the nuclear magnetic resonance (NMR) structure of recombinant human PrP (HuPrP) carrying the pathological V210I mutation responsible for familial Creutzfeldt-Jakob disease. The NMR structure of HuPrP(V210I) determined at pH 7.2 shows the same overall fold as the previously determined structure of HuPrP(V210I) at pH 5.5. It consists of a disordered N-terminal tail (residues 90-124) and a globular C-terminal domain (residues 125-231) comprising three α-helices and a short antiparallel β-sheet. Detailed comparison of three-dimensional structures of HuPrP(V210I) at pH 7.2 and 5.5 revealed significant local structural differences, with the most prominent pH-related structural variations clustered in the α(2)-α(3) interhelical region, at the interface of the β(1)-α(1) loop, in helices α(1) and α(3), and in the β(2)-α(2) loop region. The detailed analysis of interactions among secondary structure elements suggests a higher degree of structural ordering of HuPrP(V210I) under neutral-pH conditions, thus implying that spontaneous misfolding of PrP(C) may occur under acidic-pH conditions in endosomal compartments.
- Slovenian NMR Centre, National Institute of Chemistry, Hajdrihova 19, SI-1000 Ljubljana, Slovenia.
Organizational Affiliation: