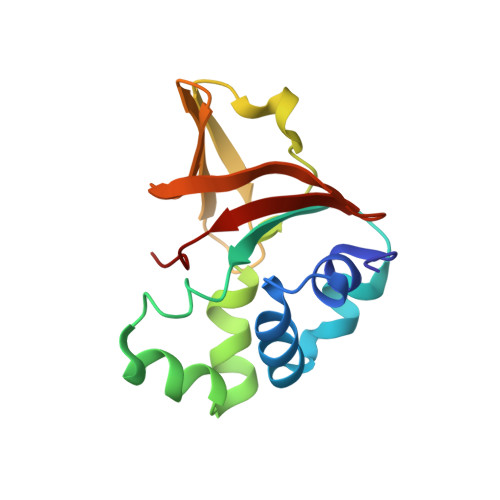
Mutation in transforming growth factor beta induced protein associated with granular corneal dystrophy type 1 reduces the proteolytic susceptibility through local structural stabilization.
Underhaug, J., Kolds, H., Runager, K., Nielsen, J.T., Srensen, C.S., Kristensen, T., Otzen, D.E., Karring, H., Malmendal, A., Schitt, B., Enghild, J.J., Nielsen, N.C.(2013) Biochim Biophys Acta 1834: 2812-2822
- PubMed: 24129074 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.bbapap.2013.10.008
- Primary Citation Related Structures:
2LTB, 2LTC - PubMed Abstract:
Hereditary mutations in the transforming growth factor beta induced (TGFBI) gene cause phenotypically distinct corneal dystrophies characterized by protein deposition in cornea. We show here that the Arg555Trp mutant of the fourth fasciclin 1 (FAS1-4) domain of the protein (TGFBIp/keratoepithelin/βig-h3), associated with granular corneal dystrophy type 1, is significantly less susceptible to proteolysis by thermolysin and trypsin than the WT domain. High-resolution liquid-state NMR of the WT and Arg555Trp mutant FAS1-4 domains revealed very similar structures except for the region around position 555. The Arg555Trp substitution causes Trp555 to be buried in an otherwise empty hydrophobic cavity of the FAS1-4 domain. The first thermolysin cleavage in the core of the FAS1-4 domain occurs on the N-terminal side of Leu558 adjacent to the Arg555 mutation. MD simulations indicated that the C-terminal end of helix α3' containing this cleavage site is less flexible in the mutant domain, explaining the observed proteolytic resistance. This structural change also alters the electrostatic properties, which may explain increased propensity of the mutant to aggregate in vitro with 2,2,2-trifluoroethanol. Based on our results we propose that the Arg555Trp mutation disrupts the normal degradation/turnover of corneal TGFBIp, leading to accumulation and increased propensity to aggregate through electrostatic interactions.
- Center for Insoluble Protein Structures (inSPIN), Interdisciplinary Nanoscience Center (iNANO) and Department of Chemistry, Aarhus University, Gustav Wieds Vej 14, DK-8000 Aarhus C, Denmark; Department of Biomedicine, University of Bergen, Jonas Lies vei 91, NO-5009 Bergen, Norway.
Organizational Affiliation: