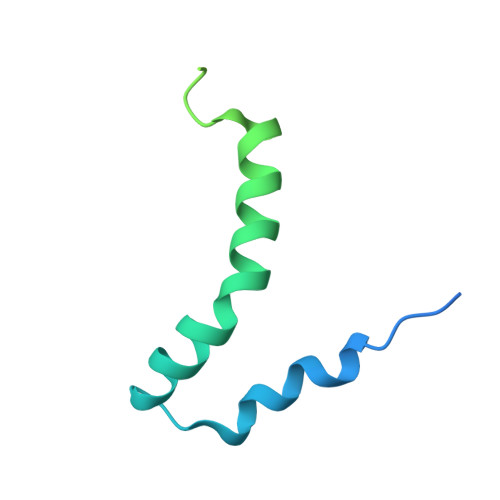
The amyloid precursor protein has a flexible transmembrane domain and binds cholesterol.
Barrett, P.J., Song, Y., Van Horn, W.D., Hustedt, E.J., Schafer, J.M., Hadziselimovic, A., Beel, A.J., Sanders, C.R.(2012) Science 336: 1168-1171
- PubMed: 22654059 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1126/science.1219988
- Primary Citation Related Structures:
2LP1 - PubMed Abstract:
C99 is the transmembrane carboxyl-terminal domain of the amyloid precursor protein that is cleaved by γ-secretase to release the amyloid-β polypeptides, which are associated with Alzheimer's disease. Nuclear magnetic resonance and electron paramagnetic resonance spectroscopy show that the extracellular amino terminus of C99 includes a surface-embedded "N-helix" followed by a short "N-loop" connecting to the transmembrane domain (TMD). The TMD is a flexibly curved α helix, making it well suited for processive cleavage by γ-secretase. Titration of C99 reveals a binding site for cholesterol, providing mechanistic insight into how cholesterol promotes amyloidogenesis. Membrane-buried GXXXG motifs (G, Gly; X, any amino acid), which have an established role in oligomerization, were also shown to play a key role in cholesterol binding. The structure and cholesterol binding properties of C99 may aid in the design of Alzheimer's therapeutics.
- Department of Biochemistry, Center for Structural Biology and Institute of Chemical Biology, Vanderbilt University School of Medicine, Nashville, TN 37232 USA.
Organizational Affiliation: