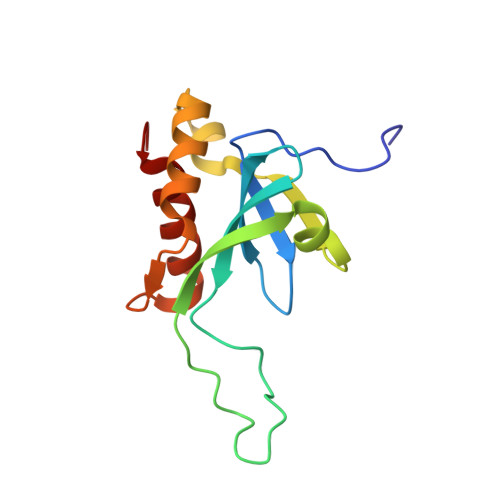
(1)H, (13)C, and (15)N resonance assignments of the N-terminal domain of human TIG3
Wang, L., Yu, W., Ren, X., Lin, J., Jin, C., Xia, B.(2012) Biomol NMR Assign 6: 201-203
- PubMed: 22290676 Search on PubMed
- DOI: https://doi.org/10.1007/s12104-012-9357-2
- Primary Citation Related Structures:
2LKT - PubMed Abstract:
Human TIG3 protein is a member of H-REV107 protein family which belongs to the type II tumor suppressor family. TIG3 can induce apoptosis in cancer cells, and it also possesses Ca(2+)-independent phospholipase A(1/2) activity. The NMR assignments of the N-terminal domain of TIG3 are essential for its solution structure determination.
- Beijing Nuclear Magnetic Resonance Center, Peking University, Beijing 100871, People’s Republic of China.
Organizational Affiliation: