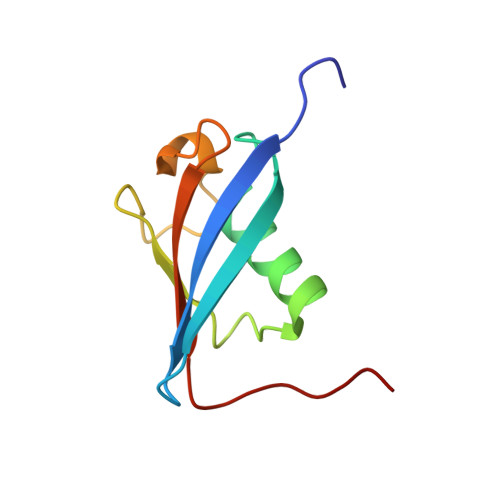
Solution structure of the E3 ligase HOIL-1 Ubl domain.
Beasley, S.A., Safadi, S.S., Barber, K.R., Shaw, G.S.(2012) Protein Sci 21: 1085-1092
- PubMed: 22517668 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/pro.2080
- Primary Citation Related Structures:
2LGY - PubMed Abstract:
The E3 ligases HOIL-1 and parkin are each comprised of an N-terminal ubiquitin-like (Ubl) domain followed by a zinc-binding region and C-terminal RING-In-between-RING-RING domains. These two proteins, involved in the ubiquitin-mediated degradation pathway, are the only two known E3 ligases to share this type of multidomain architecture. Further, the Ubl domain of both HOIL-1 and parkin has been shown to interact with the S5a subunit of the 26S proteasome. The solution structure of the HOIL-1 Ubl domain was solved using NMR spectroscopy to compare it with that of parkin to determine the structural elements responsible for S5a intermolecular interactions. The final ensemble of 20 structures had a β-grasp Ubl-fold with an overall backbone RMSD of 0.59 ± 0.10 Å in the structured regions between I55 and L131. HOIL-1 had a unique extension of both β1 and β2 sheets compared to parkin and other Ubl domains, a result of a four-residue insertion in this region. A similar 15-residue hydrophobic core in the HOIL-1 Ubl domain resulted in a comparable stability to the parkin Ubl, but significantly lower than that observed for ubiquitin. A comparison with parkin and other Ubl domains indicates that HOIL-1 likely uses a conserved hydrophobic patch (W58, V102, Y127, Y129) found on the β1 face, the β3-β4 loop and β5, as well as a C-terminal basic residue (R134) to recruit the S5a subunit as part of the ubiquitin-mediated proteolysis pathway.
- Department of Biochemistry, University of Western Ontario, London, Ontario N6A 5C1, Canada.
Organizational Affiliation: