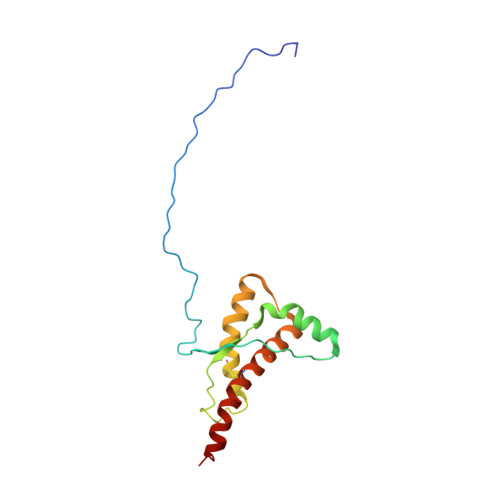
Structural basis for the protective effect of the human prion protein carrying the dominant-negative E219K polymorphism.
Biljan, I., Giachin, G., Ilc, G., Zhukov, I., Plavec, J., Legname, G.(2012) Biochem J 446: 243-251
- PubMed: 22676969 Search on PubMed
- DOI: https://doi.org/10.1042/BJ20111940
- Primary Citation Related Structures:
2LFT, 2LSB - PubMed Abstract:
The most common form of prion disease in humans is sCJD (sporadic Creutzfeldt-Jakob disease). The naturally occurring E219K polymorphism in the HuPrP (human prion protein) is considered to protect against sCJD. To gain insight into the structural basis of its protective influence we have determined the NMR structure of recombinant HuPrP (residues 90-231) carrying the E219K polymorphism. The structure of the HuPrP(E219K) protein consists of a disordered N-terminal tail (residues 90-124) and a well-structured C-terminal segment (residues 125-231) containing three α-helices and two short antiparallel β-strands. Comparison of NMR structures of the wild-type and HuPrPs with pathological mutations under identical experimental conditions revealed that, although the global architecture of the protein remains intact, replacement of Glu²¹⁹ with a lysine residue introduces significant local structural changes. The structural findings of the present study suggest that the protective influence of the E219K polymorphism is due to the alteration of surface charge distribution, in addition to subtle structural rearrangements localized within the epitopes critical for prion conversion.
- Slovenian NMR Centre, National Institute of Chemistry, Hajdrihova 19, SI-1000 Ljubljana, Slovenia.
Organizational Affiliation: