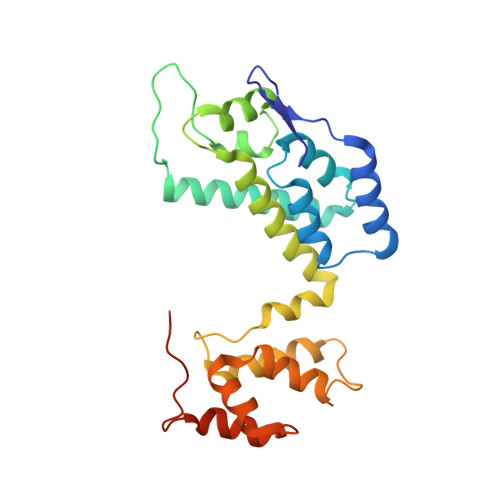
Structure of a Monomeric Mutant of the HIV-1 Capsid Protein.
Shin, R., Tzou, Y.M., Krishna, N.R.(2011) Biochemistry 50: 9457-9467
- PubMed: 21995733 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/bi2011493
- Primary Citation Related Structures:
2LF4 - PubMed Abstract:
The capsid protein (CA) of HIV-1 plays a significant role in the assembly of the immature virion and is the critical building block of its mature capsid. Thus, there has been significant interest in the CA protein as a target in the design of inhibitors of early and late stage events in the HIV-1 replication cycle. However, because of its inherent flexibility from the interdomain linker and the monomer-dimer equilibrium in solution, the HIV-1 wild-type CA monomer has defied structural determinations by X-ray crystallography and nuclear magnetic resonance spectroscopy. Here we report the detailed solution structure of full-length HIV-1 CA using a monomeric mutant that, though noninfective, preserves many of the critical properties of the wild-type protein. The structure shows independently folded N-terminal (NTD) and C-terminal domains (CTD) joined by a flexible linker. The CTD shows some differences from that of the dimeric wild-type CTD structures. This study provides insights into the molecular mechanism of the wild-type CA dimerization critical for capsid assembly. The monomeric mutant allows investigation of interactions of CA with human cellular proteins exploited by HIV-1, directly in solution without the complications associated with the monomer-dimer equilibrium of the wild-type protein. This structure also permits the design of inhibitors directed at a novel target, viz., interdomain flexibility, as well as inhibitors that target multiple interdomain interactions critical for assembly and interactions of CA with host cellular proteins that play significant roles within the replication cycle of HIV-1.
- Comprehensive Cancer Center, University of Alabama at Birmingham, Birmingham, Alabama 35294, United States.
Organizational Affiliation: