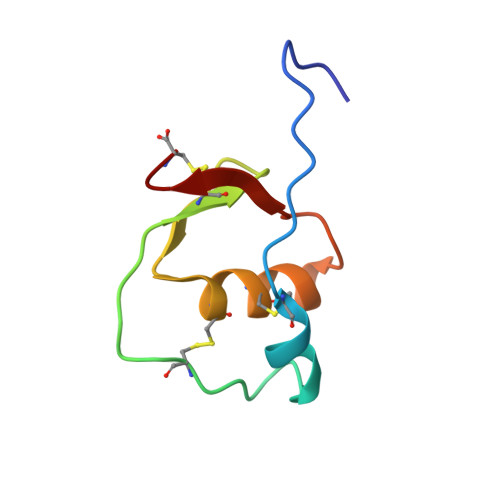
NMR structure note: human esophageal cancer-related gene 2
Feng, Y., Geng, Y., Zhou, T., Wang, J.(2012) J Biomol NMR 53: 65-70
- PubMed: 22528291 Search on PubMed
- DOI: https://doi.org/10.1007/s10858-012-9622-9
- Primary Citation Related Structures:
2LEO - National Laboratory of Biomacromolecules, Institute of Biophysics, Chinese Academy of Sciences, 15 Datun Road, Beijing 100101, China.
Organizational Affiliation: