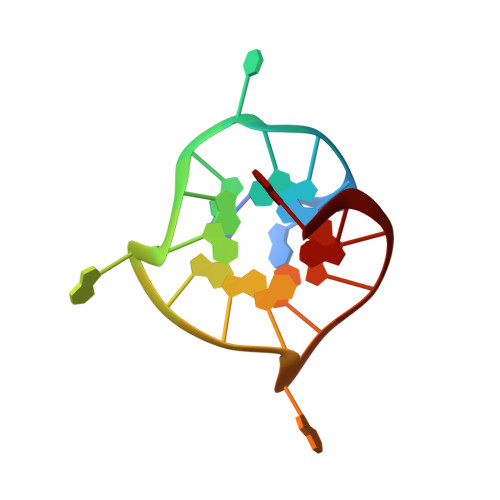
Unique Structural Features of Interconverting Monomeric and Dimeric G-Quadruplexes Adopted by a Sequence from the Intron of the N-myc Gene.
Trajkovski, M., Webba da Silva, M., Plavec, J.(2012) J Am Chem Soc 134: 4132-4141
- PubMed: 22303871 Search on PubMed
- DOI: https://doi.org/10.1021/ja208483v
- Primary Citation Related Structures:
2LED, 2LEE - PubMed Abstract:
A multidimensional heteronuclear NMR study has demonstrated that a guanine-rich DNA oligonucleotide originating from the N-myc gene folds into G-quadruplex structures in the presence of K(+), NH(4)(+), and Na(+) ions. A monomeric G-quadruplex formed in K(+) ion containing solution exhibits three G-quartets and flexible propeller-type loops. The 3D structure with three single nucleotide loops represents a missing element in structures of parallel G-quadruplexes. The structural features together with the high temperature stability are suggestive of the specific biological role of G-quadruplex formation within the intron of the N-myc gene. An increase in K(+) ion and oligonucleotide concentrations resulted in transformation of the monomeric G-quadruplex into a dimeric form. The dimeric G-quadruplex exhibits six stacked G-quartets, parallel strand orientations, and propeller-type loops. A link between the third and the fourth G-quartets consists of two adenine residues that are flipped out to facilitate consecutive stacking of six G-quartets.
- Slovenian NMR Center, National Institute of Chemistry, Hajdrihova 19, SI-1000 Ljubljana, Slovenia.
Organizational Affiliation: