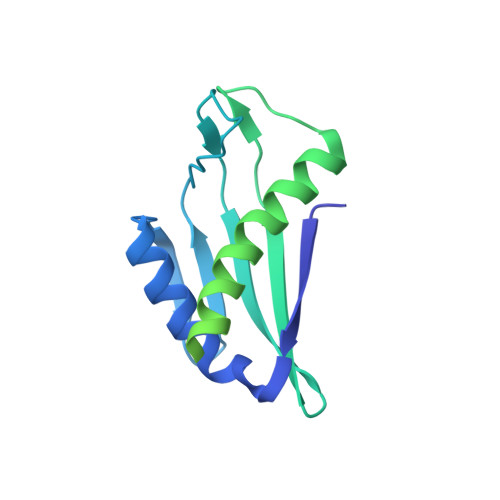
Structure of the BamC Two-Domain Protein Obtained by Rosetta with a Limited NMR Data Set.
Warner, L.R., Varga, K., Lange, O.F., Baker, S.L., Baker, D., Sousa, M.C., Pardi, A.(2011) J Mol Biology 411: 83-95
- PubMed: 21624375 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jmb.2011.05.022
- Primary Citation Related Structures:
2LAE, 2LAF - PubMed Abstract:
The CS-RDC-NOE Rosetta program was used to generate the solution structure of a 27-kDa fragment of the Escherichia coli BamC protein from a limited set of NMR data. The BamC protein is a component of the essential five-protein β-barrel assembly machine in E. coli. The first 100 residues in BamC were disordered in solution. The Rosetta calculations showed that BamC₁₀₁₋₃₄₄ forms two well-defined domains connected by an ~18-residue linker, where the relative orientation of the domains was not defined. Both domains adopt a helix-grip fold previously observed in the Bet v 1 superfamily. ¹⁵N relaxation data indicated a high degree of conformational flexibility for the linker connecting the N-terminal domain and the C-terminal domain in BamC. The results here show that CS-RDC-NOE Rosetta is robust and has a high tolerance for misassigned nuclear Overhauser effect restraints, greatly simplifying NMR structure determinations.
- Department of Chemistry and Biochemistry, University of Colorado, Boulder, Boulder, CO 80309, USA.
Organizational Affiliation: