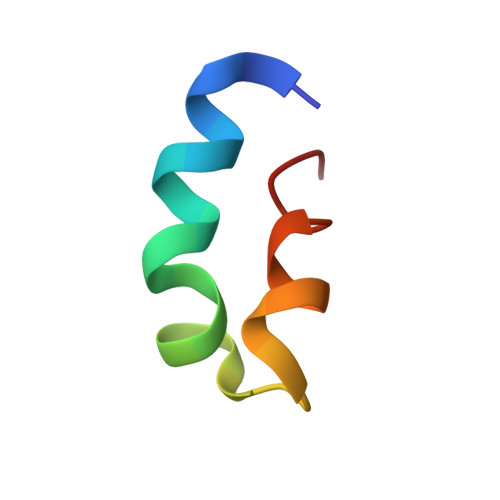
The 3D structure of thuricin CD, a two-component bacteriocin with cysteine sulfur to alpha-carbon cross-links.
Sit, C.S., McKay, R.T., Hill, C., Ross, R.P., Vederas, J.C.(2011) J Am Chem Soc 133: 7680-7683
- PubMed: 21526839 Search on PubMed
- DOI: https://doi.org/10.1021/ja201802f
- Primary Citation Related Structures:
2L9X, 2LA0 - PubMed Abstract:
Thuricin CD is an antimicrobial factor that consists of two peptides, Trn-α and Trn-β, that exhibit synergistic activity against drug resistant strains of Clostridium difficile. Trn-α and Trn-β each possess three sulfur to α-carbon thioether bridges for which the stereochemistry is unknown. This report presents the three-dimensional solution structures of Trn-α and Trn-β. Structure calculations were performed for the eight possible stereoisomers of each peptide based on the same NMR data. The structure of the stereoisomer that best fit the experimental data was chosen as the representative structure for each peptide. It was determined that Trn-α has L-stereochemistry at Ser21 (α-R), L-stereochemistry at Thr25 (α-R), and D-stereochemistry at Thr28 (α-S) (an LLD isomer). Trn-β was also found to be the LLD isomer, with L-stereochemistry at Thr21 (α-R), L-stereochemistry at Ala25 (α-R), and D-stereochemistry at Tyr28 (α-S).
- Department of Chemistry, University of Alberta, Edmonton, Alberta, Canada T6G 2G2.
Organizational Affiliation: