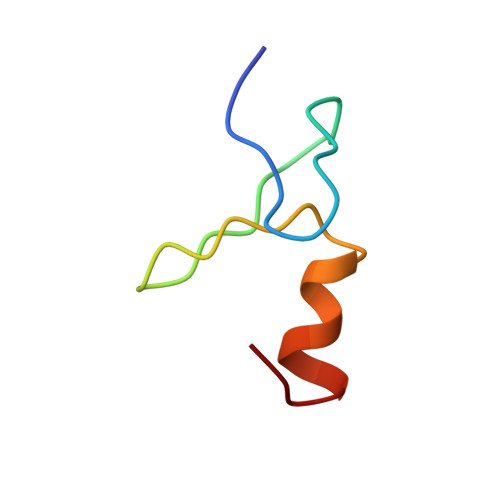
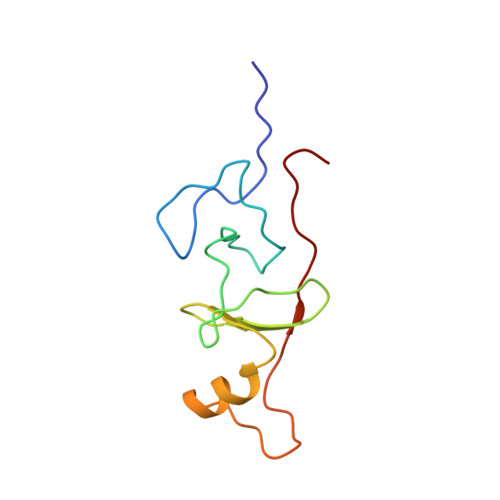
Structural basis of simultaneous recruitment of the transcriptional regulators LMO2 and FOG1/ZFPM1 by the transcription factor GATA1
Wilkinson-White, L., Gamsjaeger, R., Dastmalchi, S., Wienert, B., Stokes, P.H., Crossley, M., Mackay, J.P., Matthews, J.M.(2011) Proc Natl Acad Sci U S A 108: 14443-14448
- PubMed: 21844373 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1105898108
- Primary Citation Related Structures:
2L6Y, 2L6Z - PubMed Abstract:
The control of red blood cell and megakaryocyte development by the regulatory protein GATA1 is a paradigm for transcriptional regulation of gene expression in cell lineage differentiation and maturation. Most GATA1-regulated events require GATA1 to bind FOG1, and essentially all GATA1-activated genes are cooccupied by a TAL1/E2A/LMO2/LDB1 complex; however, it is not known whether FOG1 and TAL1/E2A/LMO2/LDB1 are simultaneously recruited by GATA1. Our structural data reveal that the FOG1-binding domain of GATA1, the N finger, can also directly contact LMO2 and show that, despite the small size (< 50 residues) of the GATA1 N finger, both FOG1 and LMO2 can simultaneously bind this domain. LMO2 in turn can simultaneously contact both GATA1 and the DNA-binding protein TAL1/E2A at bipartite E-box/WGATAR sites. Taken together, our data provide the first structural snapshot of multiprotein complex formation at GATA1-dependent genes and support a model in which FOG1 and TAL1/E2A/LMO2/LDB1 can cooccupy E-box/WGATAR sites to facilitate GATA1-mediated activation of gene activation.
- School of Molecular Bioscience, University of Sydney, New South Wales, 2006 Sydney, Australia.
Organizational Affiliation: