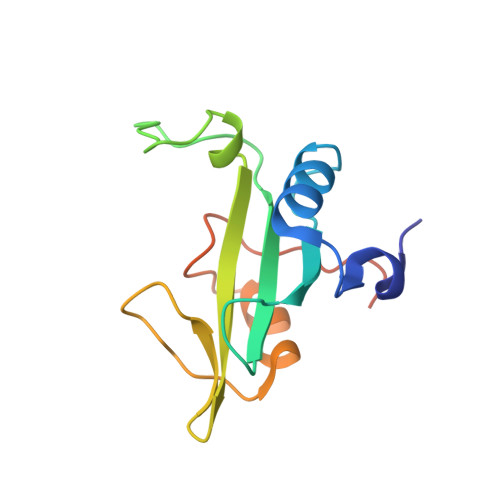
Solution structure of Tensin2 SH2 domain and its phosphotyrosine-independent interaction with DLC-1
Dai, K., Liao, S., Zhang, J., Zhang, X., Tu, X.(2011) PLoS One 6: e21965-e21965
- PubMed: 21765928 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pone.0021965
- Primary Citation Related Structures:
2L6K - PubMed Abstract:
Src homology 2 (SH2) domain is a conserved module involved in various biological processes. Tensin family member was reported to be involved in tumor suppression by interacting with DLC-1 (deleted-in-liver-cancer-1) via its SH2 domain. We explore here the important questions that what the structure of tensin2 SH2 domain is, and how it binds to DLC-1, which might reveal a novel binding mode. Tensin2 SH2 domain adopts a conserved SH2 fold that mainly consists of five β-strands flanked by two α-helices. Most SH2 domains recognize phosphorylated ligands specifically. However, tensin2 SH2 domain was identified to interact with nonphosphorylated ligand (DLC-1) as well as phosphorylated ligand. We determined the solution structure of tensin2 SH2 domain using NMR spectroscopy, and revealed the interactions between tensin2 SH2 domain and its ligands in a phosphotyrosine-independent manner.
- Hefei National Laboratory for Physical Sciences at Microscale, School of Life Sciences, University of Science and Technology of China, Hefei, Anhui, People's Republic of China.
Organizational Affiliation: