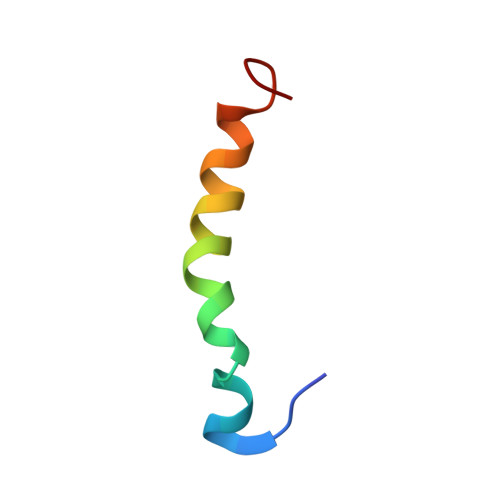
Conformational and molecular interaction studies of glucagon-like peptide-2 with its N-terminal extracellular receptor domain.
Venneti, K.C., Hewage, C.M.(2011) FEBS Lett 585: 346-352
- PubMed: 21167157 Search on PubMed
- DOI: https://doi.org/10.1016/j.febslet.2010.12.011
- Primary Citation Related Structures:
2L63, 2L64 - PubMed Abstract:
Glucagon-like peptide-2 (GLP-2) is a therapeutic target used in the treatment of short bowel syndrome. In this paper, we present the three dimensional solution structure of GLP-2 peptide determined using nuclear magnetic resonance (NMR) and molecular modelling. The GLP-2 adopts an α-helical conformation similar to that of secretin family of hormones. In order to understand the molecular details governing the ligand binding and receptor activation, macromolecular docking studies were performed between the N-terminal extracellular domain of GLP-2 receptor and the GLP-2 hormone using a data driven docking program.
- School of Biomolecular and Biomedical Science, UCD Conway Institute, University College Dublin, Dublin, Ireland.
Organizational Affiliation: