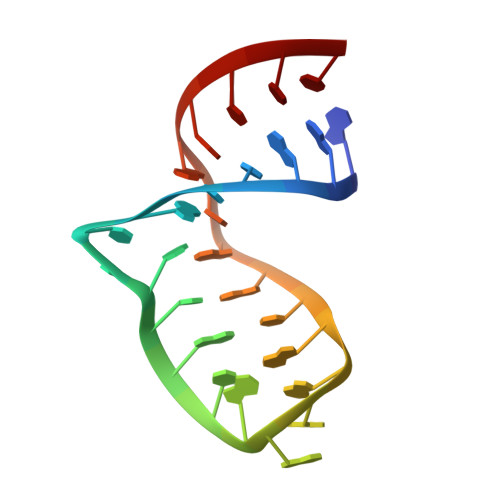
NMR structure of the A730 loop of the Neurospora VS ribozyme: insights into the formation of the active site.
Desjardins, G., Bonneau, E., Girard, N., Boisbouvier, J., Legault, P.(2011) Nucleic Acids Res 39: 4427-4437
- PubMed: 21266483 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gkq1244
- Primary Citation Related Structures:
2L5Z - PubMed Abstract:
The Neurospora VS ribozyme is a small nucleolytic ribozyme with unique primary, secondary and global tertiary structures, which displays mechanistic similarities to the hairpin ribozyme. Here, we determined the high-resolution NMR structure of a stem-loop VI fragment containing the A730 internal loop, which forms part of the active site. In the presence of magnesium ions, the A730 loop adopts a structure that is consistent with existing biochemical data and most likely reflects its conformation in the VS ribozyme prior to docking with the cleavage site internal loop. Interestingly, the A730 loop adopts an S-turn motif that is also present in loop B within the hairpin ribozyme active site. The S-turn appears necessary to expose the Watson-Crick edge of a catalytically important residue (A756) so that it can fulfill its role in catalysis. The A730 loop and the cleavage site loop of the VS ribozyme display structural similarities to internal loops found in the active site of the hairpin ribozyme. These similarities provided a rationale to build a model of the VS ribozyme active site based on the crystal structure of the hairpin ribozyme.
- Département de Biochimie, Université de Montréal, CP 6128, Succursale Centre-Ville, Montréal, Québec H3C 3J7, Canada.
Organizational Affiliation: