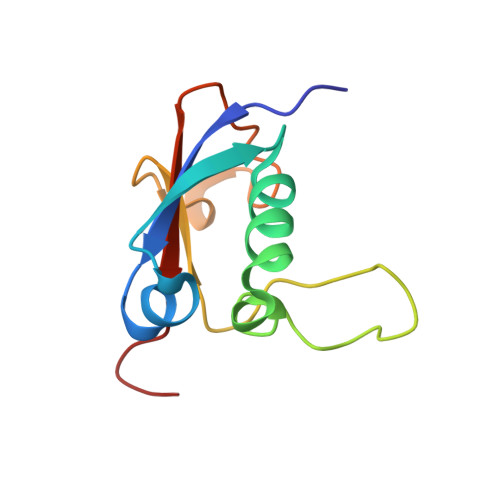
Solution structure and activation mechanism of ubiquitin-like small archaeal modifier proteins.
Ranjan, N., Damberger, F.F., Sutter, M., Allain, F.H., Weber-Ban, E.(2011) J Mol Biology 405: 1040-1055
- PubMed: 21112336 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2010.11.040
- Primary Citation Related Structures:
2L52 - PubMed Abstract:
In archaea, two ubiquitin-like small archaeal modifier protein (SAMPs) were recently shown to be conjugated to proteins in vivo. SAMPs display homology to bacterial MoaD sulfur transfer proteins and eukaryotic ubiquitin-like proteins, and they share with them the conserved C-terminal glycine-glycine motif. Here, we report the solution structure of SAMP1 from Methanosarcina acetivorans and the activation of SAMPs by an archaeal protein with homology to eukaryotic E1 enzymes. Our results show that SAMP1 possesses a β-grasp fold and that its hydrophobic and electrostatic surface features are similar to those of MoaD. M. acetivorans SAMP1 exhibits an extensive flexible surface loop between helix-2 and the third strand of the β-sheet, which contributes to an elongated surface groove that is not observed in bacterial ubiquitin homologues and many other SAMPs. We provide in vitro biochemical evidence that SAMPs are activated in an ATP-dependent manner by an E1-like enzyme that we have termed E1-like SAMP activator (ELSA). We show that activation occurs by formation of a mixed anhydride (adenylate) at the SAMP C-terminus and is detectable by SDS-PAGE and electrospray ionization mass spectrometry.
- Institute of Molecular Biology and Biophysics, Schafmattstr. 20, ETH Zürich, CH-8093 Zürich, Switzerland.
Organizational Affiliation: