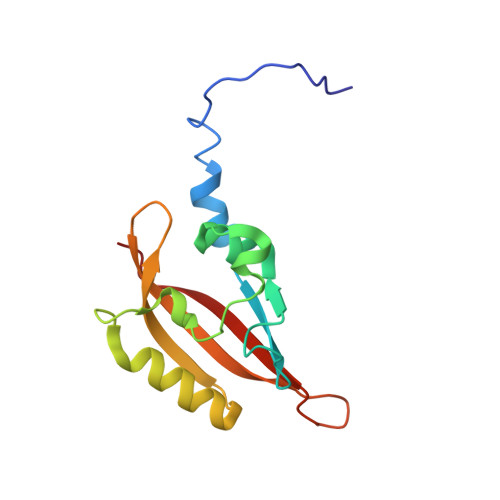
NMR solution structure of the N-terminal domain of hERG and its interaction with the S4-S5 linker.
Li, Q., Gayen, S., Chen, A.S., Huang, Q., Raida, M., Kang, C.(2010) Biochem Biophys Res Commun 403: 126-132
- PubMed: 21055387 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbrc.2010.10.132
- Primary Citation Related Structures:
2L4R - PubMed Abstract:
The human Ether-à-go-go Related Gene (hERG) potassium channel mediates the rapid delayed rectifier current (IKr) in the cardiac action potential. Mutations in the 135 amino acid residue N-terminal domain (NTD) cause channel dysfunction or mis-translocation. To study the structure of NTD, it was overexpressed and purified from Escherichia coli cells using affinity purification and gel filtration chromatography. The purified protein behaved as a monomer under purification conditions. Far- and near-UV, circular dichroism (CD) and solution nuclear magnetic resonance (NMR) studies showed that the purified protein was well-folded. The solution structure of NTD was obtained and the N-terminal residues 13-23 forming an amphipathic helix which may be important for the protein-protein or protein-membrane interactions. NMR titration experiment also demonstrated that residues from 88 to 94 in NTD are important for the molecular interaction with the peptide derived from the S4-S5 linker.
- Experimental Therapeutics Center, The Agency for Science, Technology and Research, 31 Biopolis Way Nanos, #03-01, Singapore 138669, Singapore.
Organizational Affiliation: