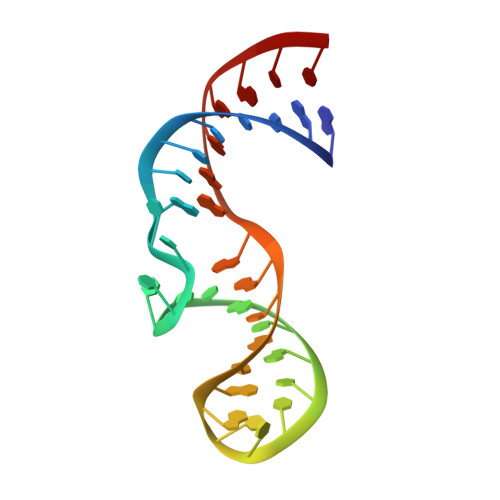
Inaugural Article: Structurally conserved five nucleotide bulge determines the overall topology of the core domain of human telomerase RNA.
Zhang, Q., Kim, N.K., Peterson, R.D., Wang, Z., Feigon, J.(2010) Proc Natl Acad Sci U S A 107: 18761-18768
- PubMed: 20966348 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1013269107
- Primary Citation Related Structures:
2L3E - PubMed Abstract:
Telomerase is a unique ribonucleoprotein complex that catalyzes the addition of telomeric DNA repeats onto the 3' ends of linear chromosomes. All vertebrate telomerase RNAs contain a catalytically essential core domain that includes the template and a pseudoknot with extended helical subdomains. Within these helical regions is an asymmetric 5-nt internal bulge loop (J2a/b) flanked by helices (P2a and P2b) that is highly conserved in its location but not sequence. NMR structure determination reveals that J2a/b forms a defined S-shape and creates an ∼90 ° bend with a surprisingly low twist (∼10 °) between the flanking helices. A search of RNA structures revealed only one other example of a 5-nt bulge, from hepatitis C virus internal ribosome entry site, with a different sequence but the same structure. J2a/b is intrinsically flexible but the interhelical motions across the loop are remarkably restricted. Nucleotide substitutions in J2a/b that affect the bend angle, direction, and interhelical dynamics are correlated with telomerase activity. Based on the structures of P2ab (J2a/b and flanking helices), the conserved region of the pseudoknot (P2b/P3, previously determined) and the remaining helical segment (P2a.1-J2a.1 refined using residual dipolar couplings and the modeling program MC-Sym) we have calculated an NMR-based model of the full-length pseudoknot. The model and dynamics analysis show that J2a/b serves as a dominant structural and dynamical element in defining the overall topology of the core domain, and suggest that interhelical motions in P2ab facilitate nucleotide addition along the template and template translocation.
- Department of Chemistry and Biochemistry, Molecular Biology Institute, PO Box 951569, University of California, Los Angeles, CA 90095-1569, USA.
Organizational Affiliation: