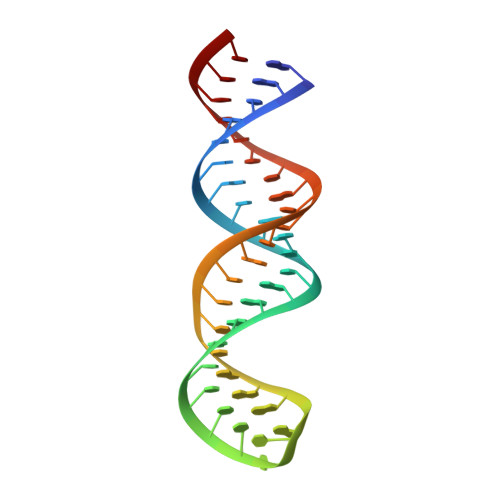
The Solution Structure of the ADAR2 dsRBM-RNA Complex Reveals a Sequence-Specific Readout of the Minor Groove.
Stefl, R., Oberstrass, F.C., Hood, J.L., Jourdan, M., Zimmermann, M., Skrisovska, L., Maris, C., Peng, L., Hofr, C., Emeson, R.B., Allain, F.H.(2010) Cell 143: 225-237
- PubMed: 20946981
- DOI: https://doi.org/10.1016/j.cell.2010.09.026
- Primary Citation Related Structures:
2L2J, 2L2K, 2L3C, 2L3J - PubMed Abstract:
Sequence-dependent recognition of dsDNA-binding proteins is well understood, yet sequence-specific recognition of dsRNA by proteins remains largely unknown, despite their importance in RNA maturation pathways. Adenosine deaminases that act on RNA (ADARs) recode genomic information by the site-selective deamination of adenosine. Here, we report the solution structure of the ADAR2 double-stranded RNA-binding motifs (dsRBMs) bound to a stem-loop pre-mRNA encoding the R/G editing site of GluR-2. The structure provides a molecular basis for how dsRBMs recognize the shape, and also more surprisingly, the sequence of the dsRNA. The unexpected direct readout of the RNA primary sequence by dsRBMs is achieved via the minor groove of the dsRNA and this recognition is critical for both editing and binding affinity at the R/G site of GluR-2. More generally, our findings suggest a solution to the sequence-specific paradox faced by many dsRBM-containing proteins that are involved in post-transcriptional regulation of gene expression.
- Institute of Molecular Biology and Biophysics, ETH Zurich, CH-8093 Zürich, Switzerland.
Organizational Affiliation: