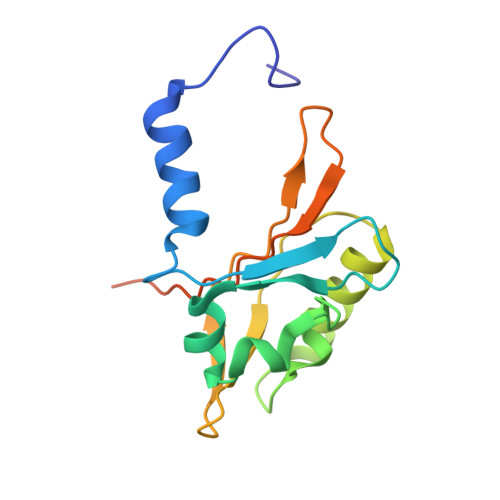
Mechanistic insight into human ether-a-go-go-related gene (hERG) K+ channel deactivation gating from the solution structure of the EAG domain.
Muskett, F.W., Thouta, S., Thomson, S.J., Bowen, A., Stansfeld, P.J., Mitcheson, J.S.(2011) J Biological Chem 286: 6184-6191
- PubMed: 21135103
- DOI: https://doi.org/10.1074/jbc.M110.199364
- Primary Citation Related Structures:
2L1M - PubMed Abstract:
Human ether-à-go-go-related gene (hERG) K(+) channels have a critical role in cardiac repolarization. hERG channels close (deactivate) very slowly, and this is vital for regulating the time course and amplitude of repolarizing current during the cardiac action potential. Accelerated deactivation is one mechanism by which inherited mutations cause long QT syndrome and potentially lethal arrhythmias. hERG deactivation is highly dependent upon an intact EAG domain (the first 135 amino acids of the N terminus). Importantly, deletion of residues 2-26 accelerates deactivation to a similar extent as removing the entire EAG domain. These and other experiments suggest the first 26 residues (NT1-26) contain structural elements required to slow deactivation by stabilizing the open conformation of the pore. Residues 26-135 form a Per-Arnt-Sim domain, but a structure for NT1-26 has not been forthcoming, and little is known about its site of interaction on the channel. In this study, we present an NMR structure for the entire EAG domain, which reveals that NT1-26 is structurally independent from the Per-Arnt-Sim domain and contains a stable amphipathic helix with one face being positively charged. Mutagenesis and electrophysiological studies indicate that neutralizing basic residues and breaking the amphipathic helix dramatically accelerate deactivation. Furthermore, scanning mutagenesis and molecular modeling studies of the cyclic nucleotide binding domain suggest that negatively charged patches on its cytoplasmic surface form an interface with the NT1-26 domain. We propose a model in which NT1-26 obstructs gating motions of the cyclic nucleotide binding domain to allosterically stabilize the open conformation of the pore.
- Department of Biochemistry, Henry Wellcome Building, University of Leicester, Lancaster Road, Leicester LE1 9HN, United Kingdom. fwm1@le.ac.uk
Organizational Affiliation: