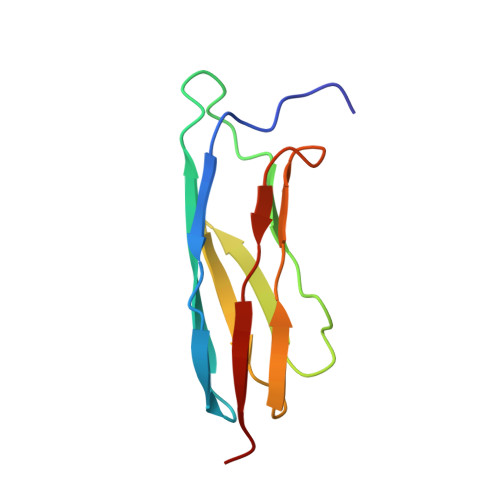
The Solution Structure of the C-Terminal Ig-like Domain of the Bacteriophage l Tail Tube Protein.
Pell, L.G., Gasmi-Seabrook, G.M., Morais, M., Neudecker, P., Kanelis, V., Bona, D., Donaldson, L.W., Edwards, A.M., Howell, P.L., Davidson, A.R., Maxwell, K.L.(2010) J Mol Biology 403: 468-479
- PubMed: 20826161 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2010.08.044
- Primary Citation Related Structures:
2L04 - PubMed Abstract:
Immunoglobulin (Ig)-like domains are found frequently on the surface of tailed double-stranded DNA bacteriophages, yet their functional role remains obscure. Here, we have investigated the structure and function of the C-terminal Ig-like domain of gpV (gpV(C)), the tail tube protein of phage λ. This domain has been predicted through sequence similarity to be a member of the bacterial Ig-like domain 2 (Big_2) family, which is composed of more than 1300 phage and bacterial sequences. Using trypsin proteolysis, we have delineated the boundaries of gpV(C) and have shown that its removal reduces the biological activity of gpV by 100-fold; thus providing a definitive demonstration of a functional role for this domain. Determination of the solution structure of gpV(C) by NMR spectroscopy showed that it adopts a canonical Ig-like fold of the I-set class. This represents the first structure of a phage-encoded Ig-like domain and only the second structure of a Big_2 domain. Structural and sequence comparisons indicate that the gpV(C) structure is more representative of both the phage-encoded Big_2 domains and Big_2 domains in general than the other available Big_2 structure. Bioinformatics analyses have identified two conserved clusters of residues on the surface of gpV(C) that may be important in mediating the function of this domain.
- Department of Biochemistry, Faculty of Medicine, University of Toronto, Medical Sciences Building, Toronto, ON, Canada.
Organizational Affiliation: