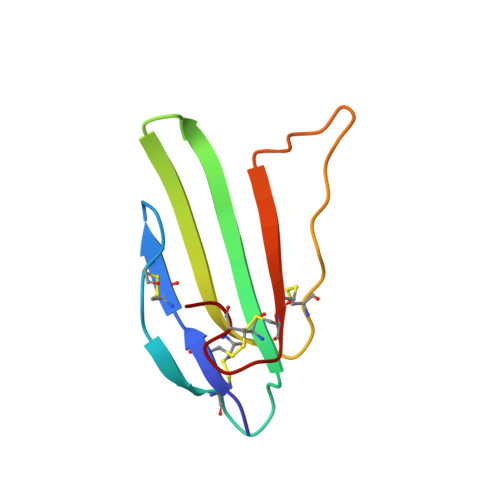
NMR structure and action on nicotinic acetylcholine receptors of water-soluble domain of human LYNX1
Lyukmanova, E.N., Shenkarev, Z.O., Shulepko, M.A., Mineev, K.S., D'Hoedt, D., Kasheverov, I.E., Filkin, S.Y., Krivolapova, A.P., Janickova, H., Dolezal, V., Dolgikh, D.A., Arseniev, A.S., Bertrand, D., Tsetlin, V.I., Kirpichnikov, M.P.(2011) J Biological Chem 286: 10618-10627
- PubMed: 21252236 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M110.189100
- Primary Citation Related Structures:
2L03 - PubMed Abstract:
Discovery of proteins expressed in the central nervous system sharing the three-finger structure with snake α-neurotoxins provoked much interest to their role in brain functions. Prototoxin LYNX1, having homology both to Ly6 proteins and three-finger neurotoxins, is the first identified member of this family membrane-tethered by a GPI anchor, which considerably complicates in vitro studies. We report for the first time the NMR spatial structure for the water-soluble domain of human LYNX1 lacking a GPI anchor (ws-LYNX1) and its concentration-dependent activity on nicotinic acetylcholine receptors (nAChRs). At 5-30 μM, ws-LYNX1 competed with (125)I-α-bungarotoxin for binding to the acetylcholine-binding proteins (AChBPs) and to Torpedo nAChR. Exposure of Xenopus oocytes expressing α7 nAChRs to 1 μM ws-LYNX1 enhanced the response to acetylcholine, but no effect was detected on α4β2 and α3β2 nAChRs. Increasing ws-LYNX1 concentration to 10 μM caused a modest inhibition of these three nAChR subtypes. A common feature for ws-LYNX1 and LYNX1 is a decrease of nAChR sensitivity to high concentrations of acetylcholine. NMR and functional analysis both demonstrate that ws-LYNX1 is an appropriate model to shed light on the mechanism of LYNX1 action. Computer modeling, based on ws-LYNX1 NMR structure and AChBP x-ray structure, revealed a possible mode of ws-LYNX1 binding.
- Shemyakin-Ovchinnikov Institute of Bioorganic Chemistry, Russian Academy of Sciences, 16/10 Miklukho-Maklaya Street, 117997 Moscow, Russia.
Organizational Affiliation: