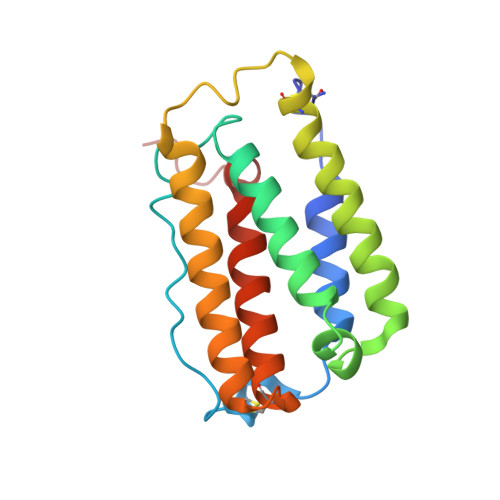
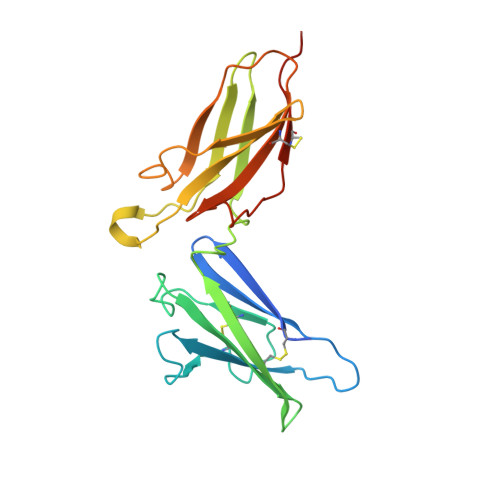
Intermolecular interactions in a 44 kDa interferon-receptor complex detected by asymmetric reverse-protonation and two-dimensional NOESY
Nudelman, I., Akabayov, S.R., Schnur, E., Biron, Z., Levy, R., Xu, Y., Yang, D., Anglister, J.(2010) Biochemistry 49: 5117-5133
- PubMed: 20496919 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/bi100041f
- Primary Citation Related Structures:
2KZ1 - PubMed Abstract:
Type I interferons (IFNs) make up a family of homologous helical cytokines initiating strong antiviral and antiproliferative activity. All type I IFNs bind to a common cell surface receptor consisting of two subunits, IFNAR1 and IFNAR2, associating upon binding of interferon. We studied intermolecular interactions between IFNAR2-EC and IFNalpha2 using asymmetric reverse-protonation of the different complex components and two-dimensional homonuclear NOESY. This new approach revealed with an excellent signal-to-noise ratio 24 new intermolecular NOEs between the two molecules despite the low concentration of the complex (0.25 mM) and its high molecular mass (44 kDa). Sequential and side chain assignment of IFNAR2-EC and IFNalpha2 in their binary complex helped assign the intermolecular NOEs to the corresponding protons. A docking model of the IFNAR2-EC-IFNalpha2 complex was calculated on the basis of the intermolecular interactions found in this study as well as four double mutant cycle constraints, previously observed NOEs between a single pair of residues and the NMR mapping of the binding sites on IFNAR2-EC and IFNalpha2. Our docking model doubles the buried surface area of the previous model and significantly increases the number of intermolecular hydrogen bonds, salt bridges, and van der Waals interactions. Furthermore, our model reveals the participation of several new regions in the binding site such as the N-terminus and A helix of IFNalpha2 and the C domain of IFNAR2-EC. As a result of these additions, the orientation of IFNAR2-EC relative to IFNalpha2 has changed by 30 degrees in comparison with a previously calculated model that was based on NMR mapping of the binding sites and double mutant cycle constraints. In addition, the new model strongly supports the recently proposed allosteric changes in IFNalpha2 upon binding of IFNAR1-EC to the binary IFNalpha2-IFNAR2-EC complex.
- Department of Structural Biology, Weizmann Institute of Science, Rehovot 76100, Israel.
Organizational Affiliation: