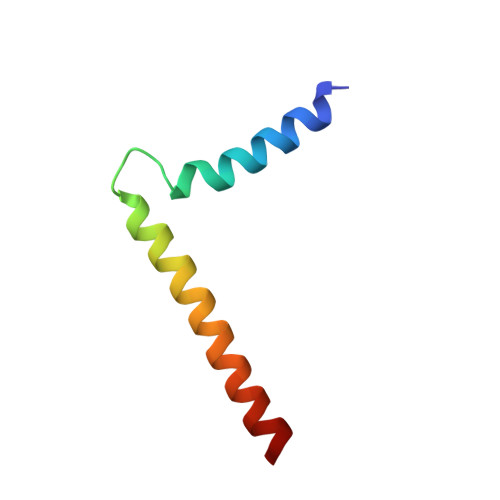
Structural topology of phospholamban pentamer in lipid bilayers by a hybrid solution and solid-state NMR method.
Verardi, R., Shi, L., Traaseth, N.J., Walsh, N., Veglia, G.(2011) Proc Natl Acad Sci U S A 108: 9101-9106
- PubMed: 21576492 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1016535108
- Primary Citation Related Structures:
2KYV - PubMed Abstract:
Phospholamban (PLN) is a type II membrane protein that inhibits the sarcoplasmic reticulum Ca(2+)-ATPase (SERCA), thereby regulating calcium homeostasis in cardiac muscle. In membranes, PLN forms pentamers that have been proposed to function either as a storage for active monomers or as ion channels. Here, we report the T-state structure of pentameric PLN solved by a hybrid solution and solid-state NMR method. In lipid bilayers, PLN adopts a pinwheel topology with a narrow hydrophobic pore, which excludes ion transport. In the T state, the cytoplasmic amphipathic helices (domains Ia) are absorbed into the lipid bilayer with the transmembrane domains arranged in a left-handed coiled-coil configuration, crossing the bilayer with a tilt angle of approximately 11° with respect to the membrane normal. The tilt angle difference between the monomer and pentamer is approximately 13°, showing that intramembrane helix-helix association forces dominate over the hydrophobic mismatch, driving the overall topology of the transmembrane assembly. Our data reveal that both topology and function of PLN are shaped by the interactions with lipids, which fine-tune the regulation of SERCA.
- Department of Biochemistry, University of Minnesota, Minneapolis, MN 55455, USA.
Organizational Affiliation: