Solution structure of MAST2-PDZ complexed with the C-terminus of PTEN
Terrien, E., Wolff, N., Cordier, F., Simenel, C., Lafon, M., Prehaud, C., Delepierre, M.To be published.
Experimental Data Snapshot
wwPDB Validation 3D Report Full Report
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Microtubule-associated serine/threonine-protein kinase 2 | 96 | Homo sapiens | Mutation(s): 0 EC: 2.7.11.1 | 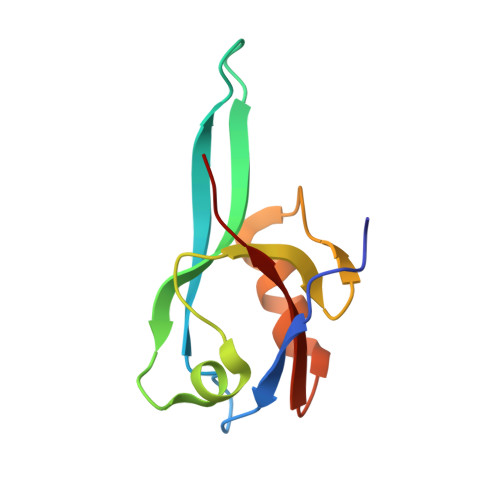 | |
UniProt & NIH Common Fund Data Resources | |||||
PHAROS: Q6P0Q8 GTEx: ENSG00000086015 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | Q6P0Q8 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| C-terminus of Phosphatidylinositol-3,4,5-trisphosphate 3-phosphatase and dual-specificity protein phosphatase PTEN | 13 | N/A | Mutation(s): 0 EC: 3.1.3.67 (UniProt), 3.1.3 (UniProt), 3.1.3.16 (UniProt), 3.1.3.48 (UniProt) | 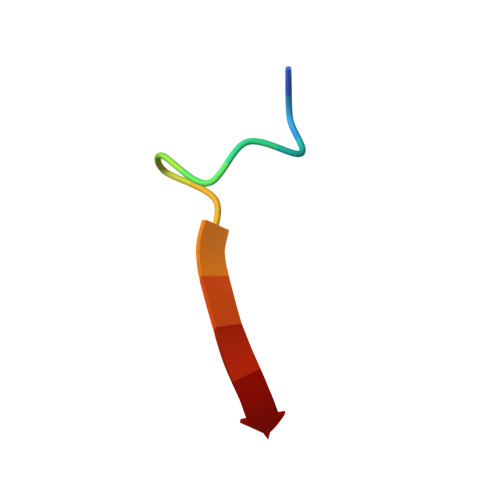 | |
UniProt & NIH Common Fund Data Resources | |||||
PHAROS: P60484 GTEx: ENSG00000171862 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P60484 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||