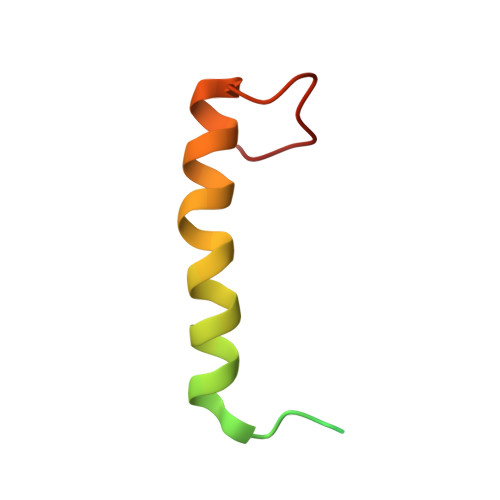
Residues within a lipid-associated segment of the PECAM-1 cytoplasmic domain are susceptible to inducible, sequential phosphorylation.
Paddock, C., Lytle, B.L., Peterson, F.C., Holyst, T., Newman, P.J., Volkman, B.F., Newman, D.K.(2011) Blood 117: 6012-6023
- PubMed: 21464369 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1182/blood-2010-11-317867
- Primary Citation Related Structures:
2KY5 - PubMed Abstract:
Immunoreceptor tyrosine-based inhibitory motif (ITIM)-containing receptors inhibit cellular responsiveness to immunoreceptor tyrosine-based activation motif (ITAM)-linked receptors. Although tyrosine phosphorylation is central to the initiation of both inhibitory ITIM and stimulatory ITAM signaling, the events that regulate receptor phosphorylation are incompletely understood. Previous studies have shown that ITAM tyrosines engage in structure-inducing interactions with the plasma membrane that must be relieved for phosphorylation to occur. Whether ITIM phosphorylation is similarly regulated and the mechanisms responsible for release from plasma membrane interactions to enable phosphorylation, however, have not been defined. PECAM-1 is a dual ITIM-containing receptor that inhibits ITAM-dependent responses in hematopoietic cells. We found that the PECAM-1 cytoplasmic domain is unstructured in an aqueous environment but adopts an α-helical conformation within a localized region on interaction with lipid vesicles that mimic the plasma membrane. The lipid-interacting segment contains the C-terminal ITIM tyrosine and a serine residue that undergo activation-dependent phosphorylation. The N-terminal ITIM is excluded from the lipid-interacting segment, and its phosphorylation is secondary to phosphorylation of the membrane-interacting C-terminal ITIM. On the basis of these findings, we propose a novel model for regulation of inhibitory signaling by ITIM-containing receptors that relies on reversible plasma membrane interactions and sequential ITIM phosphorylation.
- Blood Research Institute, BloodCenter of Wisconsin and Department of Biochemistry, Medical College of Wisconsin, Milwaukee, WI 53226, USA.
Organizational Affiliation: