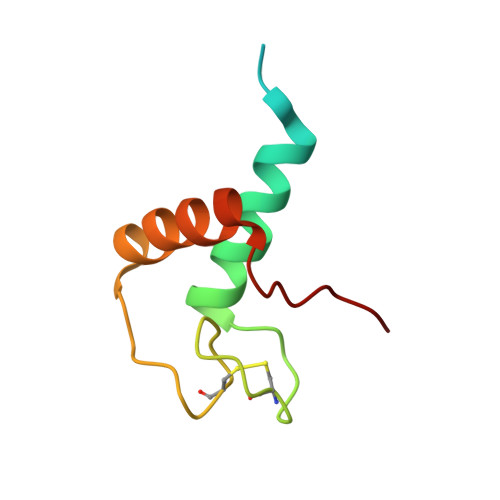
Zn-binding AZUL domain of human ubiquitin protein ligase Ube3A.
Lemak, A., Yee, A., Bezsonova, I., Dhe-Paganon, S., Arrowsmith, C.H.(2011) J Biomol NMR 51: 185-190
- PubMed: 21947926 Search on PubMed
- DOI: https://doi.org/10.1007/s10858-011-9552-y
- Primary Citation Related Structures:
2KR1 - PubMed Abstract:
Ube3A (also referred to as E6AP for E6 Associated Protein) is a E3 ubiquitin-protein ligase implicated in the development of Angelman syndrome by controlling degradation of synaptic protein Arc and oncogenic papilloma virus infection by controlling degradation of p53. This article describe the solution NMR structure of the conserved N-terminal domain of human Ube3A (residues 24-87) that contains two residues (Cys44 and Arg62) found to be mutated in patients with Angelman syndrome. The structure of this domain adopts a novel Zn-binding fold we called AZUL (Amino-terminal Zn-finger of Ube3a Ligase). The AZUL domain has a helix-loop-helix architecture with a Zn ion coordinated by four Cys residues arranged in Cys-X(4)-Cys-X(4)-Cys-X(28)-Cys motif. Three of the Zn-bound residues are located in a 23-residue long and well structured loop that connects two α-helicies.
- Ontario Cancer Institute, Campbell Family Cancer Research Institute and Department of Medical Biophysics, University of Toronto, and Northeast Structural Genomics Consortium, 101 College Street, Toronto, ON M5G 1L7, Canada.
Organizational Affiliation: