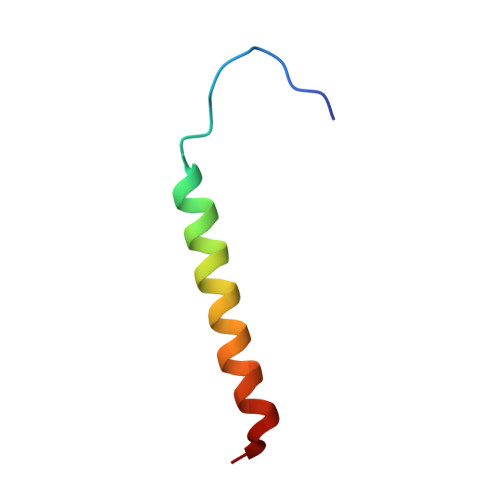
Dimeric structure of the transmembrane domain of glycophorin a in lipidic and detergent environments.
Mineev, K.S., Bocharov, E.V., Volynsky, P.E., Goncharuk, M.V., Tkach, E.N., Ermolyuk, Y.S., Schulga, A.A., Chupin, V.V., Maslennikov, I.V., Efremov, R.G., Arseniev, A.S.(2011) Acta Naturae 3: 90-98
- PubMed: 22649687 Search on PubMedSearch on PubMed Central
- Primary Citation Related Structures:
2KPE, 2KPF - PubMed Abstract:
Specific interactions between transmembrane α-helices, to a large extent, determine the biological function of integral membrane proteins upon normal development and in pathological states of an organism. Various membrane-like media, partially those mimicking the conditions of multicomponent biological membranes, are used to study the structural and thermodynamic features that define the character of oligomerization of transmembrane helical segments. The choice of the composition of the membrane-mimicking medium is conducted in an effort to obtain a biologically relevant conformation of the protein complex and a sample that would be stable enough to allow to perform a series of long-term experiments with its use. In the present work, heteronuclear NMR spectroscopy and molecular dynamics simulations were used to demonstrate that the two most widely used media (detergent DPC micelles and lipid DMPC/DHPC bicelles) enable to perform structural studies of the specific interactions between transmembrane α-helices by the example of dimerizing the transmembrane domain of the bitopic protein glycophorin A. However, a number of peculiarities place lipid bicelles closer to natural lipid bilayers in terms of their physical properties.
- Shemyakin and Ovchinnikov Institute of Bioorganic Chemistry, Russian Academy of Sciences.
Organizational Affiliation: