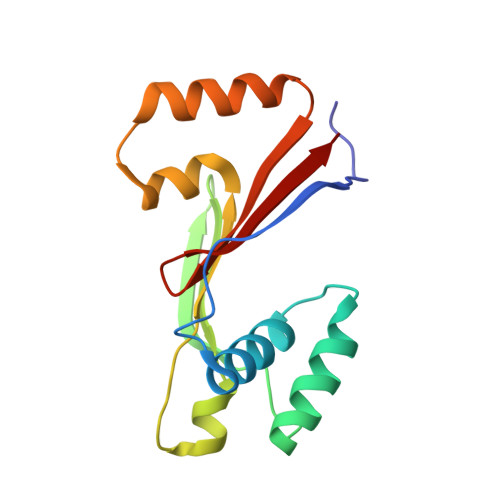
Difference in stability of the N-domain underlies distinct intracellular properties of the E1064A and H1069Q mutants of copper-transporting ATPase ATP7B.
Dmitriev, O.Y., Bhattacharjee, A., Nokhrin, S., Uhlemann, E.M., Lutsenko, S.(2011) J Biological Chem 286: 16355-16362
- PubMed: 21398519 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M110.198101
- Primary Citation Related Structures:
2KOY - PubMed Abstract:
Wilson disease (WD) is a disorder of copper metabolism caused by mutations in the Cu-transporting ATPase ATP7B. WD is characterized by significant phenotypic variability, the molecular basis of which is poorly understood. The E1064A mutation in the N-domain of ATP7B was previously shown to disrupt ATP binding. We have now determined, by NMR, the structure of the N-domain containing this mutation and compared properties of E1064A and H1069Q, another mutant with impaired ATP binding. The E1064A mutation does not change the overall fold of the N-domain. However, the position of the α1,α2-helical hairpin (α-HH) that houses Glu(1064) and His(1069) is altered. The α-HH movement produces a more open structure compared with the wild-type ATP-bound form and misaligns ATP coordinating residues, thus explaining complete loss of ATP binding. In the cell, neither the stability nor targeting of ATP7B-E1064A to the trans-Golgi network differs significantly from the wild type. This is in a contrast to the H1069Q mutation within the same α-HH, which greatly destabilizes protein both in vitro and in cells. The difference between two mutants can be linked to a lower stability of the α-HH in the H1069Q variant at the physiological temperature. We conclude that the structural stability of the N-domain rather than the loss of ATP binding plays a defining role in the ability of ATP7B to reach the trans-Golgi network, thus contributing to phenotypic variability in WD.
- Department of Biochemistry, University of Saskatchewan, Saskatoon, Saskatchewan, Canada. oleg.dmitriev@usask.ca
Organizational Affiliation: