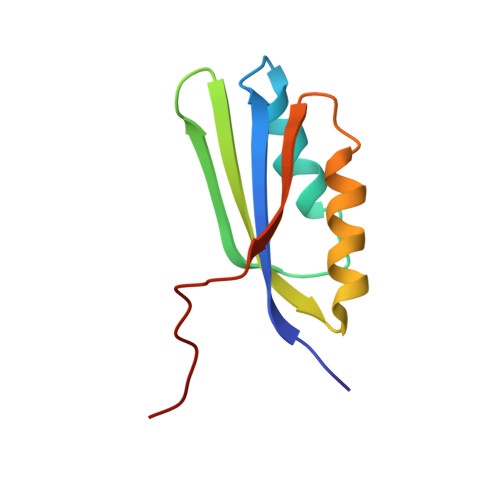
Solution NMR structure of the ACT domain from GTP pyrophosphokinase of Chlorobium tepidum
Eletsky, A., Garcia, E., Wang, H., Ciccosanti, C., Jiang, M., Nair, R., Rost, B., Acton, T.B., Xiao, R., Everett, J.K., Lee, H., Prestegard, J., Montelione, G.T., Szyperski, T.To be published.