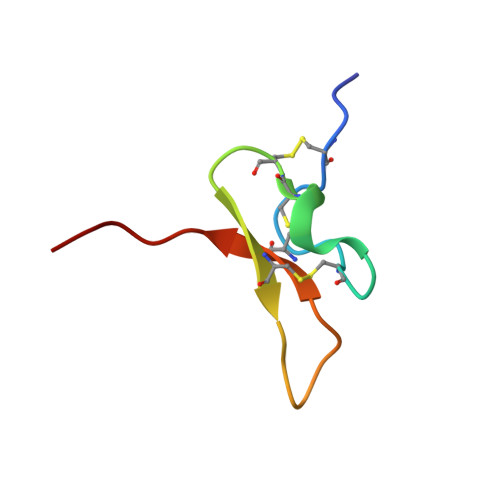
A dynamic pharmacophore drives the interaction between Psalmotoxin-1 and the putative drug target acid-sensing ion channel 1a.
Saez, N.J., Mobli, M., Bieri, M., Chassagnon, I.R., Malde, A.K., Gamsjaeger, R., Mark, A.E., Gooley, P.R., Rash, L.D., King, G.F.(2011) Mol Pharmacol 80: 796-808
- PubMed: 21825095 Search on PubMed
- DOI: https://doi.org/10.1124/mol.111.072207
- Primary Citation Related Structures:
2KNI - PubMed Abstract:
Acid-sensing ion channel 1a (ASIC1a) is a primary acid sensor in the peripheral and central nervous system. It has been implicated as a novel therapeutic target for a broad range of pathophysiological conditions including pain, ischemic stroke, depression, and autoimmune diseases such as multiple sclerosis. The only known selective blocker of ASIC1a is π-TRTX-Pc1a (PcTx1), a disulfide-rich 40-residue peptide isolated from spider venom. π-TRTX-Pc1a is an effective analgesic in rodent models of acute pain and it provides neuroprotection in a mouse model of ischemic stroke. Thus, understanding the molecular basis of the π-TRTX-Pc1a-ASIC1a interaction should facilitate development of therapeutically useful ASIC1a blockers. We therefore developed an efficient bacterial expression system to produce a panel of π-TRTX-Pc1a mutants for probing structure-activity relationships as well as isotopically labeled toxin for determination of its solution structure and dynamics. We demonstrate that the toxin pharmacophore resides in a β-hairpin loop that was revealed to be mobile over a wide range of time scales using molecular dynamics simulations in combination with NMR spin relaxation and relaxation dispersion measurements. The toxin-receptor interaction was modeled by in silico docking of the toxin structure onto a homology model of rat ASIC1a in a restraints-driven approach that was designed to take account of the dynamics of the toxin pharmacophore and the consequent remodeling of side-chain conformations upon receptor binding. The resulting model reveals new insights into the mechanism of action of π-TRTX-Pc1a and provides an experimentally validated template for the rational design of therapeutically useful π-TRTX-Pc1a mimetics.
- Institute for Molecular Bioscience, University of Queensland, 306 Carmody Road, St Lucia, QLD 4072, Australia.
Organizational Affiliation: