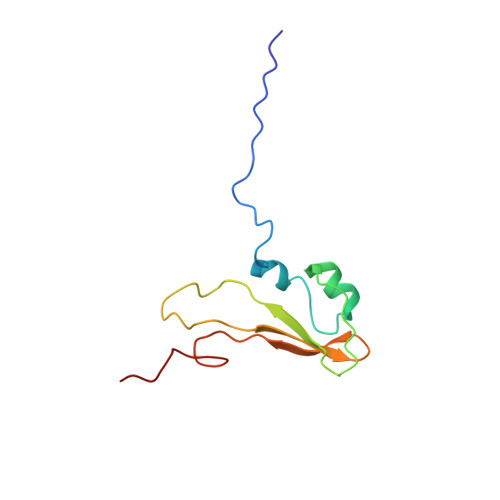
Structure and function of BamE within the outer membrane and the beta-barrel assembly machine.
Knowles, T.J., Browning, D.F., Jeeves, M., Maderbocus, R., Rajesh, S., Sridhar, P., Manoli, E., Emery, D., Sommer, U., Spencer, A., Leyton, D.L., Squire, D., Chaudhuri, R.R., Viant, M.R., Cunningham, A.F., Henderson, I.R., Overduin, M.(2011) EMBO Rep 12: 123-128
- PubMed: 21212804 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/embor.2010.202
- Primary Citation Related Structures:
2KM7 - PubMed Abstract:
Insertion of folded proteins into the outer membrane of Gram-negative bacteria is mediated by the essential β-barrel assembly machine (Bam). Here, we report the native structure and mechanism of a core component of this complex, BamE, and show that it is exclusively monomeric in its native environment of the periplasm, but is able to adopt a distinct dimeric conformation in the cytoplasm. BamE is shown to bind specifically to phosphatidylglycerol, and comprehensive mutagenesis and interaction studies have mapped key determinants for complex binding, outer membrane integrity and cell viability, as well as revealing the role of BamE within the Bam complex.
- School of Cancer Sciences, University of Birmingham, Edgbaston, Birmingham B15 2TT, UK.
Organizational Affiliation: