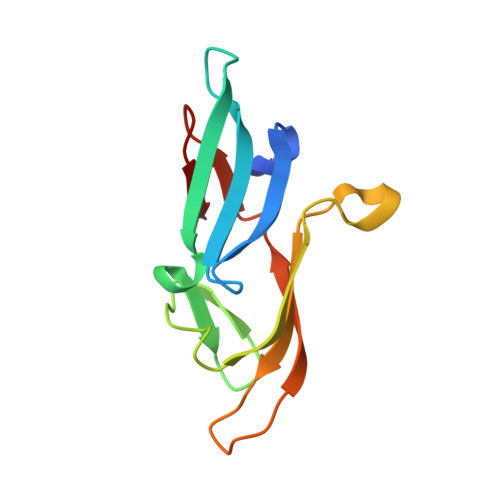
The evolutionarily conserved family of cyanovirin-N homologs: structures and carbohydrate specificity.
Koharudin, L.M., Viscomi, A.R., Jee, J.G., Ottonello, S., Gronenborn, A.M.(2008) Structure 16: 570-584
- PubMed: 18400178 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2008.01.015
- Primary Citation Related Structures:
2JZJ, 2JZK, 2JZL, 2KJL - PubMed Abstract:
Solution structures for three members of the recently discovered cyanovirin-N (CV-N) homolog family of lectins have been determined. Cyanovirin-N homologs (CVNHs) from Tuber borchii, Ceratopteris richardii, and Neurospora crassa, representing each of the three phylogenetic groups, were selected. All proteins exhibit the same fold, and the overall structures resemble that of the founding member of the family, CV-N, albeit with noteworthy differences in loop conformation and detailed local structure. Since no data are available regarding the proteins' function or their natural ligands, extensive carbohydrate-binding studies were conducted. We delineated ligand-binding sites on all three proteins by nuclear magnetic resonance and identified which sugars interact by array screening. The number and location of binding sites vary for the three proteins, and different ligand specificities exist. Potential physiological roles for two family members, TbCVNH and NcCVNH, were probed in nutrition deprivation experiments that suggest a possible involvement of these proteins in lifestyle-related responses.
- Department of Structural Biology, School of Medicine, University of Pittsburgh, Biomedical Science Tower 3, 3501 Fifth Avenue, Pittsburgh, PA 15260, USA.
Organizational Affiliation: