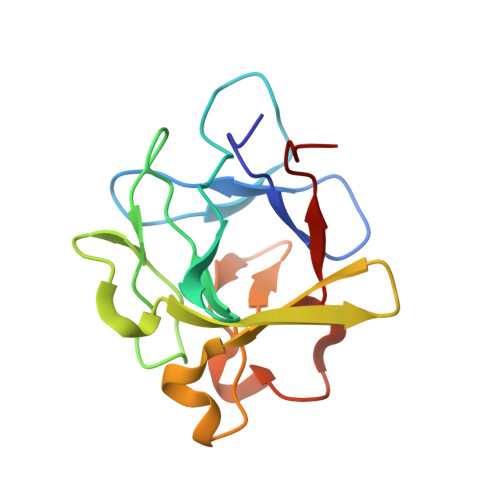
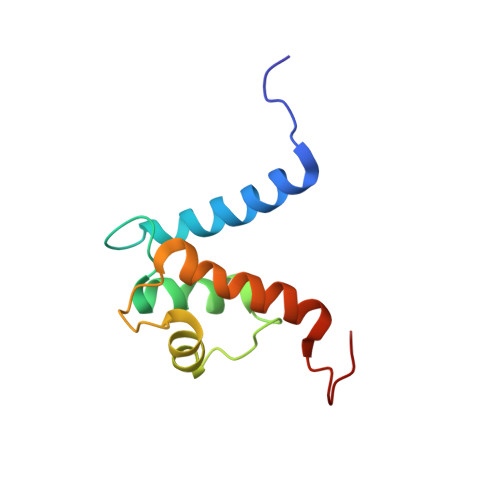
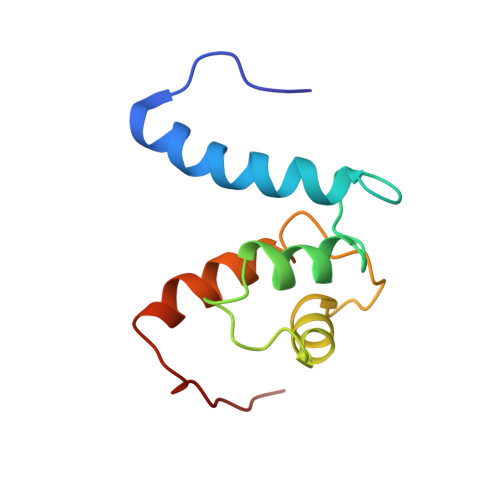
The heterohexameric complex structure, a component in the non-classical pathway for fibroblast growth factor 1 (FGF1) secretion.
Mohan, S.K., Rani, S.G., Yu, C.(2010) J Biological Chem 285: 15464-15475
- PubMed: 20220137 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M109.066357
- Primary Citation Related Structures:
2KI4, 2KI6 - PubMed Abstract:
Fibroblast growth factors (FGFs) are key regulators of cell proliferation, tumor-induced angiogenesis, and migration. FGFs are essential for early embryonic development, organ formation, and angiogenesis. FGF1 also plays an important role in inflammation, wound healing, and restenosis. The biological effects of FGF1 are mediated through the activation of the four transmembrane phosphotyrosine kinase fibroblast growth factor receptors in the presence of heparin sulfate proteoglycans and, therefore, require the release of the protein into the extracellular space. FGF1 is exported through a non-classical release pathway involving the formation of a specific multiprotein complex. The protein constituents of this complex include FGF1, S100A13, and the p40 form of synaptotagmin 1 (Syt1). Because FGF1 plays an important role in tumor formation, it is clear that preventing the formation of the multiprotein complex would be an effective strategy to inhibit a wide range of cancers. To understand the molecular events in the FGF1 release pathway, we studied the FGF1-S100A13 tetrameric and FGF1-S100A13-C2A hexameric complex structures, which are both complexes possibly formed during the non-classical pathway of FGF1 release.
- Department of Chemistry, National Tsing Hua University, Hsinchu 30013, Taiwan.
Organizational Affiliation: