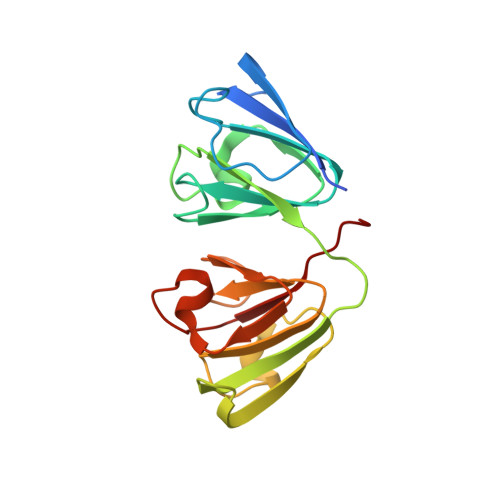
The structure of the cataract-causing P23T mutant of human gammaD-crystallin exhibits distinctive local conformational and dynamic changes.
Jung, J., Byeon, I.J., Wang, Y., King, J., Gronenborn, A.M.(2009) Biochemistry 48: 2597-2609
- PubMed: 19216553 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/bi802292q
- Primary Citation Related Structures:
2KFB - PubMed Abstract:
Crystallins are major proteins of the eye lens and essential for lens transparency. Mutations and aging of crystallins cause cataracts, the predominant cause of blindness in the world. In human gammaD-crystallin, the P23T mutant is associated with congenital cataracts. Until now, no atomic structural information has been available for this variant. Biophysical analyses of this mutant protein have revealed dramatically reduced solubility compared to that of the wild-type protein due to self-association into higher-molecular weight clusters and aggregates that retain a nativelike conformation within the monomers [Pande, A., et al. (2005) Biochemistry 44, 2491-2500]. To elucidate the structure and local conformation around the mutation site, we have determined the solution structure and characterized the protein's dynamic behavior by NMR. Although the global structure is very similar to the X-ray structure of wild-type gammaD-crystallin, pivotal local conformational and dynamic differences are caused by the threonine substitution. In particular, in the P23T mutant, the imidazole ring of His22 switches from the predominant Nepsilon2 tautomer in the wild-type protein to the Ndelta1 tautomer, and an altered motional behavior of the associated region in the protein is observed. The data support structural changes that may initiate aggregation or polymerization by the mutant protein.
- Department of Structural Biology, School of Medicine, University of Pittsburgh, Pittsburgh, Pennsylvania 15260, USA.
Organizational Affiliation: