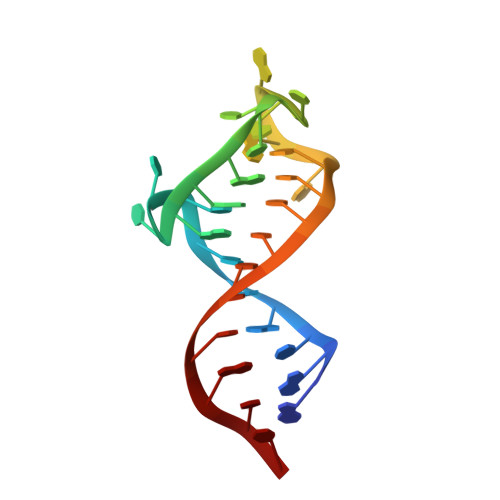
Simultaneous recognition of HIV-1 TAR RNA bulge and loop sequences by cyclic peptide mimics of Tat protein
Davidson, A., Leeper, T.C., Athanassiou, Z., Patora-Komisarska, K., Karn, J., Robinson, J.A., Varani, G.(2009) Proc Natl Acad Sci U S A 106: 11931-11936
- PubMed: 19584251 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.0900629106
- Primary Citation Related Structures:
2KDQ - PubMed Abstract:
The interaction of the HIV-1 transactivator protein Tat with its transactivation response (TAR) RNA is an essential step in viral replication and therefore an attractive target for developing antivirals with new mechanisms of action. Numerous compounds that bind to the 3-nt bulge responsible for binding Tat have been identified in the past, but none of these molecules had sufficient potency to warrant pharmaceutical development. We have discovered conformationally-constrained cyclic peptide mimetics of Tat that are specific nM inhibitors of the Tat-TAR interaction by using a structure-based approach. The lead peptides are nearly as active as the antiviral drug nevirapine against a variety of clinical isolates in human lymphocytes. The NMR structure of a peptide-RNA complex reveals that these molecules interfere with the recruitment to TAR of both Tat and the essential cellular cofactor transcription elongation factor-b (P-TEFb) by binding simultaneously at the RNA bulge and apical loop, forming an unusually deep pocket. This structure illustrates additional principles in RNA recognition: RNA-binding molecules can achieve specificity by interacting simultaneously with multiple secondary structure elements and by inducing the formation of deep binding pockets in their targets. It also provides insight into the P-TEFb binding site and a rational basis for optimizing the promising antiviral activity observed for these cyclic peptides.
- Department of Chemistry, University of Washington, Box 351700, Seattle, WA 98195, USA.
Organizational Affiliation: