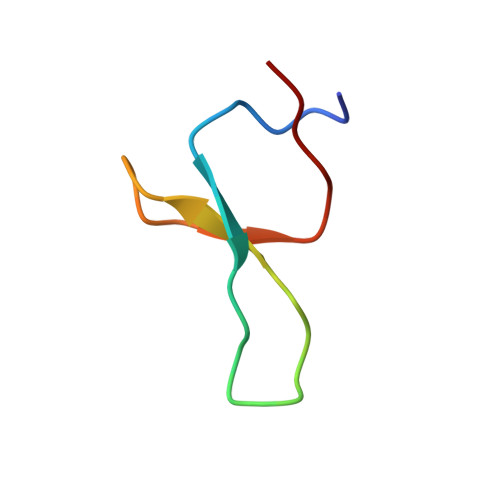
NMR solution structure of the isolated Apo Pin1 WW domain: comparison to the x-ray crystal structures of Pin1
Kowalski, J.A., Liu, K., Kelly, J.W.(2002) Biopolymers 63: 111-121
- PubMed: 11786999 Search on PubMed
- DOI: https://doi.org/10.1002/bip.10020
- Primary Citation Related Structures:
2KCF - PubMed Abstract:
The NMR solution structure of the isolated Apo Pin1 WW domain (6-39) reveals that it adopts a twisted three-stranded antiparallel beta-sheet conformation, very similar to the structure exhibited by the crystal of this domain in the context of the two domain Pin1 protein. While the B factors in the apo x-ray crystal structure indicate that loop 1 and loop 2 are conformationally well defined, the solution NMR data suggest that loop 1 is quite flexible, at least in the absence of the ligand. The NMR chemical shift and nuclear Overhauser effect pattern exhibited by the 6-39 Pin1 WW domain has proven to be diagnostic for demonstrating that single site variants of this domain adopt a normally folded structure. Knowledge of this type is critical before embarking on time-consuming kinetic and thermodynamic studies required for a detailed understanding of beta-sheet folding.
- Department of Chemistry, Skaggs Institute of Chemical Biology, The Scripps Research Institute, BCC265 10550 N. Torrey Pines Road, La Jolla, CA 92037.
Organizational Affiliation: