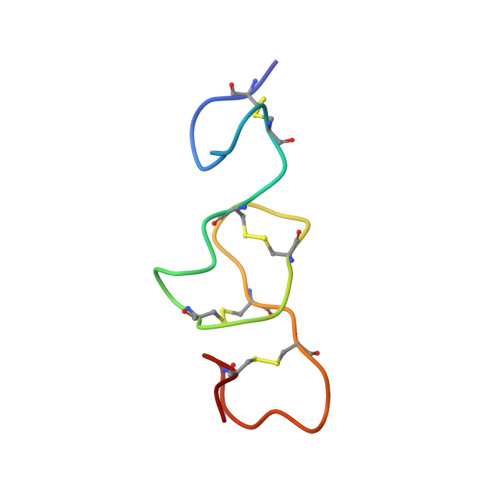
Exon 6 of human JAG1 encodes a conserved structural unit
Pintar, A., Guarnaccia, C., Dhir, S., Pongor, S.(2009) BMC Struct Biol 9: 43-43
- PubMed: 19586525 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1186/1472-6807-9-43
- Primary Citation Related Structures:
2KB9 - PubMed Abstract:
Notch signaling drives developmental processes in all metazoans. The receptor binding region of the human Notch ligand Jagged-1 is made of a DSL (Delta/Serrate/Lag-2) domain and two atypical epidermal growth factor (EGF) repeats encoded by two exons, exon 5 and 6, which are out of phase with respect to the EGF domain boundaries. We determined the 1H-NMR solution structure of the polypeptide encoded by exon 6 of JAG1 and spanning the C-terminal region of EGF1 and the entire EGF2. We show that this single, evolutionary conserved exon defines an autonomous structural unit that, despite the minimal structural context, closely matches the structure of the same region in the entire receptor binding module. In eukaryotic genomes, exon and domain boundaries usually coincide. We report a case study where this assertion does not hold, and show that the autonomously folding, structural unit is delimited by exon boundaries, rather than by predicted domain boundaries.
- International Centre for Genetic Engineering and Biotechnology, Protein Structure and Bioinformatics Group, AREA Science Park, Padriciano 99, Trieste, Italy. pintar@icgeb.org
Organizational Affiliation: