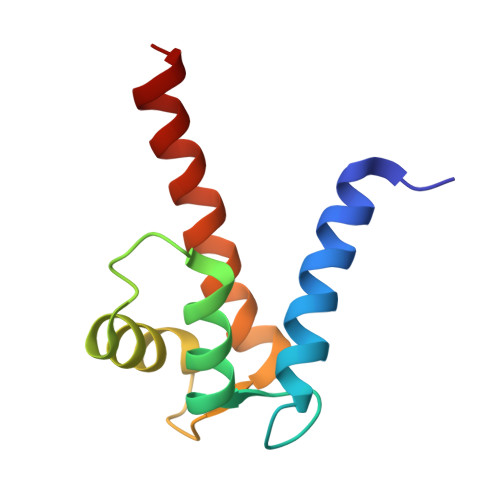
Solution structure and dynamics of S100A5 in the apo and Ca2+-bound states
Bertini, I., Das Gupta, S., Hu, X., Karavelas, T., Luchinat, C., Parigi, G., Yuan, J.(2009) J Biol Inorg Chem 14: 1097-1107
- PubMed: 19536568 Search on PubMed
- DOI: https://doi.org/10.1007/s00775-009-0553-1
- Primary Citation Related Structures:
2KAX, 2KAY - PubMed Abstract:
S100A5 is a calcium binding protein of the S100 family, with one canonical and one S100-specific EF-hand motif per subunit. Although its function is still unknown, it has recently been reported to be one of the S100 proteins able to interact with the receptor for advanced glycation end products. The homodimeric solution structures of S100A5 in both the apo and the calcium(II)-loaded forms have been obtained, and show a conformational rearrangement upon calcium binding. This rearrangement involves, in particular, the hinge loop connecting the N-terminal and the C-terminal EF-hand domains, the reorientation of helix III with respect to helix IV, as common to several S100 proteins, and the elongation of helix IV. The details of the structural changes are important because they must be related to the different functions, still largely unknown, of the different members of the S100 family. For the first time for a full-length S100 protein, relaxation measurements were performed on both the apo and the calcium-bound forms. A quite large mobility was observed in the hinge loop, which is not quenched in the calcium form. The structural differences resulting upon calcium binding change the global shape and the distribution of hydrophobic and charged residues of the S100A5 homodimer in a modest but significantly different manner with respect to the closest homologues S100A4 and S100A6.
- Magnetic Resonance Center (CERM), University of Florence, Via Luigi Sacconi 6, 50019 Sesto Fiorentino, Italy. ivanobertini@cerm.unifi.it
Organizational Affiliation: