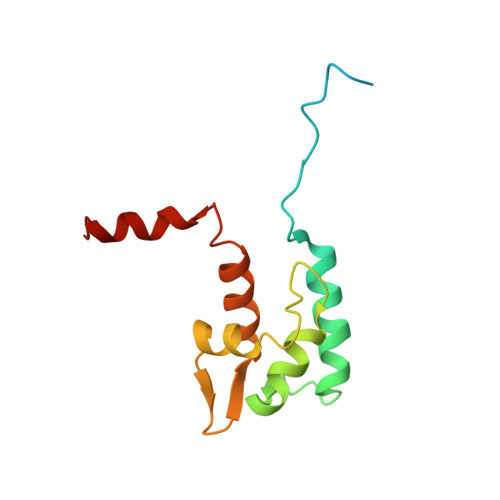
Solution structure of the NaV1.2 C-terminal EF-hand domain.
Miloushev, V.Z., Levine, J.A., Arbing, M.A., Hunt, J.F., Pitt, G.S., Palmer, A.G.(2009) J Biological Chem 284: 6446-6454
- PubMed: 19129176 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M807401200
- Primary Citation Related Structures:
2KAV - PubMed Abstract:
Voltage-gated sodium channels initiate the rapid upstroke of action potentials in many excitable tissues. Mutations within intracellular C-terminal sequences of specific channels underlie a diverse set of channelopathies, including cardiac arrhythmias and epilepsy syndromes. The three-dimensional structure of the C-terminal residues 1777-1882 of the human NaV1.2 voltage-gated sodium channel has been determined in solution by NMR spectroscopy at pH 7.4 and 290.5 K. The ordered structure extends from residues Leu-1790 to Glu-1868 and is composed of four alpha-helices separated by two short anti-parallel beta-strands; a less well defined helical region extends from residue Ser-1869 to Arg-1882, and a disordered N-terminal region encompasses residues 1777-1789. Although the structure has the overall architecture of a paired EF-hand domain, the NaV1.2 C-terminal domain does not bind Ca2+ through the canonical EF-hand loops, as evidenced by monitoring 1H,15N chemical shifts during aCa2+ titration. Backbone chemical shift resonance assignments and Ca2+ titration also were performed for the NaV1.5 (1773-1878) isoform, demonstrating similar secondary structure architecture and the absence of Ca2+ binding by the EF-hand loops. Clinically significant mutations identified in the C-terminal region of NaV1 sodium channels cluster in the helix I-IV interface and the helix II-III interhelical segment or in helices III and IV of the NaV1.2 (1777-1882) structure.
- Department of Biochemistry and Molecular Biophysics, Columbia University, New York, New York 10032-3702, USA.
Organizational Affiliation: