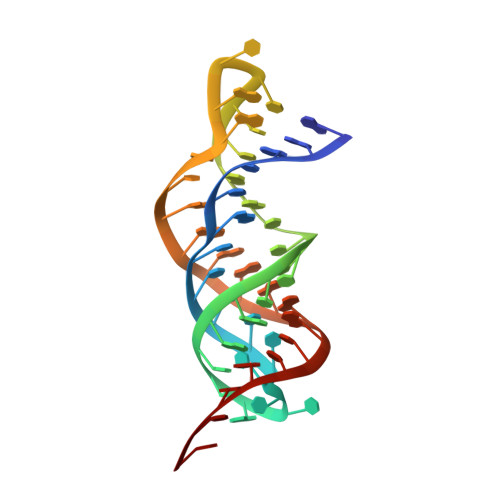
Solution Structure and Dynamics of the Wild-type Pseudoknot of Human Telomerase RNA.
Kim, N.K., Zhang, Q., Zhou, J., Theimer, C.A., Peterson, R.D., Feigon, J.(2008) J Mol Biology 384: 1249-1261
- PubMed: 18950640
- DOI: https://doi.org/10.1016/j.jmb.2008.10.005
- Primary Citation Related Structures:
2K95, 2K96 - PubMed Abstract:
Telomerase is a ribonucleoprotein complex that replicates the 3' ends of linear chromosomes by successive additions of telomere repeat DNA. The telomerase holoenzyme contains two essential components for catalysis, a telomerase reverse transcriptase (TERT) and telomerase RNA (TER). The TER includes a template for telomere repeat synthesis as well as other domains required for function. We report the solution structure of the wild-type minimal conserved human TER pseudoknot refined with an extensive set of RDCs, and a detailed analysis of the effect of the bulge U177 on pseudoknot structure, dynamics analyzed by RDC and 13C relaxation measurements, and base pair stability. The overall structure of PKWT is highly similar to the previously reported DeltaU177 pseudoknot (PKDU) that has a deletion of a conserved bulge U important for catalytic activity. For direct comparison to PKWT, the structure of PKDU was re-refined with a comparable set of RDCs. Both pseudoknots contain a catalytically essential triple helix at the junction of the two stems, including two stem 1-loop 2 minor groove triples, a junction loop 1-loop 2 Hoogsteen base pair, and stem 2-loop 1 major groove U.A-U Watson-Crick-Hoogsteen triples located directly above the bulge U177. However, there are significant differences in the stabilities of base pairs near the bulge and the dynamics of some nucleotides. The stability of the base pairs in stem 2 surrounding the bulge U177 is greatly decreased, with the result that the Watson-Crick pairs in the triple helix begin to unfold before the Hoogsteen pairs, which may affect telomerase assembly and activity. The bulge U is positioned in the minor groove on the face opposite the triple helical interactions, and sterically blocks the A176 2'OH, which has recently been proposed to have a role in catalysis. The bulge U may serve as a hinge providing backbone flexibility in this region.
- Department of Chemistry and Biochemistry, P.O. Box 951569, University of California, Los Angeles, CA 90095-1569, USA.
Organizational Affiliation: