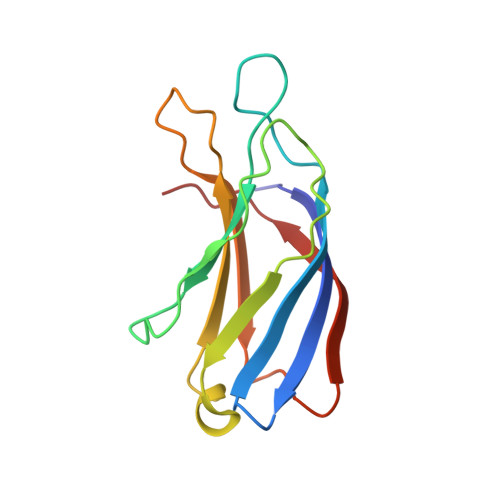
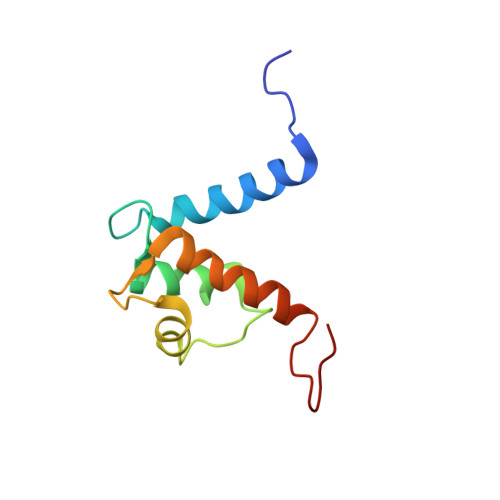
S100A13-C2A binary complex structure-a key component in the acidic fibroblast growth factor for the non-classical pathway.
Mohan, S.K., Rani, S.G., Kumar, S.M., Yu, C.(2009) Biochem Biophys Res Commun 380: 514-519
- PubMed: 19284995 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbrc.2009.01.143
- Primary Citation Related Structures:
2K8M - PubMed Abstract:
Fibroblast growth factors (FGFs) are key regulators of cell proliferation, differentiation, tumor-induced angiogenesis and migration. FGFs are essential for early embryonic development, organ formation and angiogenesis. They play important roles in tumor formation, inflammation, wound healing and restenosis. The biological effects of FGFs are mediated through the activation of the four transmembrane phosphotyrosine kinase receptors (FGFRs) in the presence of heparin sulfate proteoglycans (HSPGs) and therefore require the release of FGFs into the extracellular space. However, FGF-1 lacks the signal peptide required for the releasing of these proteins through the classical endoplasmic reticulum (ER)-Golgi secretary pathway. Maciag et al. demonstrated that FGF-1 is exported through a non-classical release pathway involving the formation of a specific multiprotein complex [M. Landriscina, R. Soldi, C. Bagala, I. Micucci, S. Bellum, F. Tarantini, I. Prudovsky, T. Maciag, S100A13 participates in the release of fibroblast growth factor 1 in response to heat shock in vitro, J. Biol. Chem. 276 (2001) 22544-22552; C.M. Carreira, T.M. LaVallee, F. Tarantini, A. Jackson, J.T. Lathrop, B. Hampton, W.H. Burgess, T. Maciag, S100A13 is involved in the regulation of fibroblast growth factor-1 and p40 synaptotagmin-1 release in vitro, J. Biol. Chem. 273 (1998) 22224-22231; T.M. LaValle, F. Tarantini, S. Gamble, C.M. Carreira, A. Jackson, T. Maciag, Synaptotagmin-1 is required for fibroblast growth factor-1 release, J. Biol. Chem. 273 (1998) 22217-22223; C. Bagalá, V. Kolev, A. Mandinova, R. Soldi, C. Mouta, I. Graziani, I, Prudovsky, T. Maciag, The alternative translation of synaptotagmin 1 mediates the non-classical release of FGF1, Biochem. Biophys. Res. Commun. 310 (2003) 1041-1047]. The protein constituents of this complex include FGF-1, S100A13 (a Ca(2+)-binding protein), and the p40 form of synaptotagmin 1 (Syt1). To understand the molecular events in the FGF-1 releasing pathway, we have studied the interactions of S100A13 with C2A by (1)H-(15)N HSQC titration and 3D-filtered NOESY experiments. We characterized the binary complex structure of S100A13-C2A by using a variety of multi-dimensional NMR experiments. This complex acts as a template for FGF-1 dimerization and multiprotein complex formation.
- Department of Chemistry, National Tsing Hua University, Hsinchu 30013, Taiwan.
Organizational Affiliation: