The human Cdc37.Hsp90 complex studied by heteronuclear NMR spectroscopy
Sreeramulu, S., Jonker, H.R.A., Langer, T., Richter, C., Lancaster, C.R., Schwalbe, H.(2009) J Biological Chem 284: 3885-3896
- PubMed: 19073599 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M806715200
- Primary Citation Related Structures:
2K5B, 2W0G - PubMed Abstract:
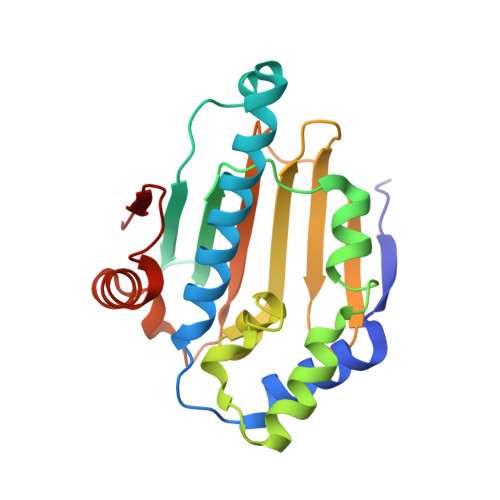
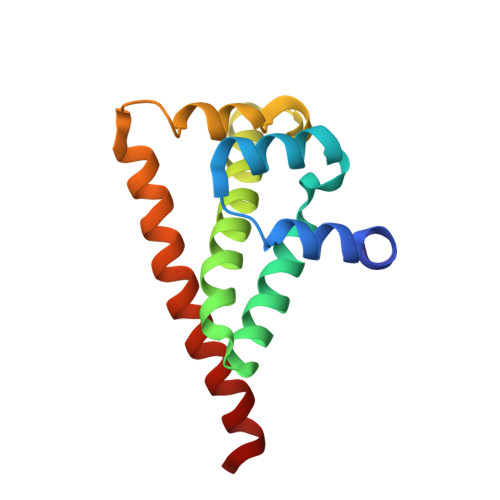
The cell division cycle protein 37 (Cdc37) and the 90-kDa heat shock protein (Hsp90) are molecular chaperones, which are crucial elements in the protein signaling pathway. The largest class of client proteins for Cdc37 and Hsp90 are protein kinases. The catalytic domains of these kinases are stabilized by Cdc37, and their proper folding and functioning is dependent on Hsp90. Here, we present the x-ray crystal structure of the 16-kDa middle domain of human Cdc37 at 1.88 angstroms resolution and the structure of this domain in complex with the 23-kDa N-terminal domain of human Hsp90 based on heteronuclear solution state NMR data and docking. Our results demonstrate that the middle domain of Cdc37 exists as a monomer. NMR and mutagenesis experiments reveal Leu-205 in Cdc37 as a key residue enabling complex formation. These findings can be very useful in the development of small molecule inhibitors against cancer.
- Institute for Organic Chemistry and Chemical Biology, Center for Biomolecular Magnetic Resonance, Johann Wolfgang Goethe-University, Max-von-Laue-Strasse 7, Frankfurt am Main D-60438, Germany.
Organizational Affiliation: