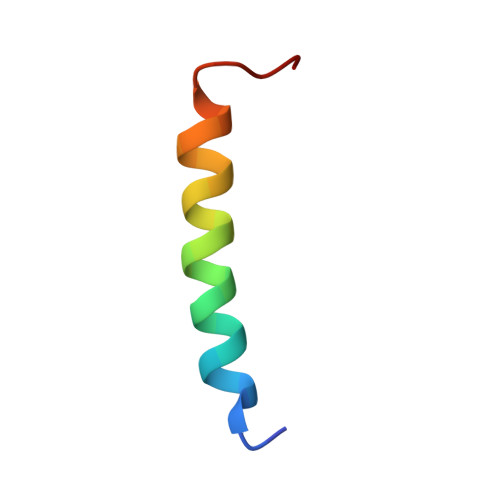
NMR structure and dynamics of the second transmembrane domain of the neuronal acetylcholine receptor beta 2 subunit
Yushmanov, V.E., Xu, Y., Tang, P.(2003) Biochemistry 42: 13058-13065
- PubMed: 14596621 Search on PubMed
- DOI: https://doi.org/10.1021/bi0350396
- Primary Citation Related Structures:
2K59 - PubMed Abstract:
Structure and backbone dynamics of a selectively [(15)N]Leu-labeled 28-residue segment of the extended second transmembrane domain (TM2e) of the human neuronal nicotinic acetylcholine receptor (nAChR) beta(2) subunit were studied by (1)H and (15)N solution-state NMR in dodecylphosphocholine micelles. The TM2e structure was determined on the basis of the nuclear Overhauser effects (NOEs) and the hydrogen bond restraints, which were inferred from the presence of H(alpha)(i)-H(N)(i+3), H(alpha)(i)-H(beta)(i+3), and H(alpha)(i)-H(N)(i+4) NOE connectivity and from the slow amide hydrogen exchange with D(2)O. The TM2e structure of the nAChR beta(2) subunit contains a helical region between T4 and K22. Backbone dynamics were calculated using the model-free approach based on the (15)N relaxation rate constants, R(1) and R(2), and on the (15)N-[(1)H] NOE. The data acquired at 9.4 and 14.1 T and calculations using different dynamic models demonstrated no conformational exchange and internal motions on the nanosecond time scale. The global tumbling time of TM2e in micelles was 14.4 +/- 0.2 ns; the NOE values were greater than 0.63 at 9.4 T, and the order parameter, S(2), was 0.83-0.96 for all (15)N-labeled leucine residues, suggesting a restricted internal motion. This is the first report of NMR structure and backbone dynamics of the second transmembrane domain of the human nAChR beta(2) subunit in a membrane-mimetic environment, providing the basis for subsequent studies of subunit interactions in the transmembrane domain complex of the neuronal nAChR.
- Department of Anesthesiology, University of Pittsburgh School of Medicine, Pittsburgh, Pennsylvania 15261, USA.
Organizational Affiliation: